SIRT1 regulates macrophage self-renewal
- PMID: 28701484
- PMCID: PMC5556267
- DOI: 10.15252/embj.201695737
SIRT1 regulates macrophage self-renewal
Abstract
Mature differentiated macrophages can self-maintain by local proliferation in tissues and can be extensively expanded in culture under specific conditions, but the mechanisms of this phenomenon remain only partially defined. Here, we show that SIRT1, an evolutionary conserved regulator of life span, positively affects macrophage self-renewal ability in vitro and in vivo Overexpression of SIRT1 during bone marrow-derived macrophage differentiation increased their proliferative capacity. Conversely, decrease of SIRT1 expression by shRNA inactivation, CRISPR/Cas9 mediated deletion and pharmacological inhibition restricted macrophage self-renewal in culture. Furthermore, pharmacological SIRT1 inhibition in vivo reduced steady state and cytokine-induced proliferation of alveolar and peritoneal macrophages. Mechanistically, SIRT1 inhibition negatively regulated G1/S transition, cell cycle progression and a network of self-renewal genes. This included inhibition of E2F1 and Myc and concomitant activation of FoxO1, SIRT1 targets mediating cell cycle progression and stress response, respectively. Our findings indicate that SIRT1 is a key regulator of macrophage self-renewal that integrates cell cycle and longevity pathways. This suggests that macrophage self-renewal might be a relevant parameter of ageing.
Keywords: cell cycle regulation; macrophage; replicative life span; self‐renewal; sirtuins.
© 2017 The Authors.
Figures
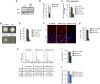
Immunoblot for SIRT1 protein comparing bone marrow‐derived wild‐type (WT BMM) and MafB/c‐Maf double knockout (Maf‐DKO) macrophages. Grb2 antibody was used as loading control.
Quantification of panel (A). Shown are Sirt1/Grb2 ratios (arbitrary units, A.U.), normalized to Grb2. Error bars indicate the standard error of the mean. Each condition was done in duplicate; data represent the pool of two independent experiments.
Quantitative PCR for the expression of SIRT1 comparing Maf‐DKO macrophages infected with indicated shRNA vectors to non‐infected Maf‐DKO and wild‐type (WT) macrophages. Shown are fold changes of the average values normalized to HPRT of two independent experiments and standard error of the mean.
Effect of SIRT1 inactivation on the colony formation potential of Maf‐DKO macrophages. Phase contrast magnification ×10. Each condition was done in duplicate; the results shown are representative of two independent experiments. Scale bars = 50 μm.
Quantification of panel (D). Data represent the pool of two independent experiments. Error bars indicate SEM.
Immunostaining for SIRT1 (red) on Maf‐DKO macrophages infected with shRNA vectors against LacZ or SIRT1. DAPI (blue) was used to stain DNA. Each condition was done in duplicate; the results shown are representative of two independent experiments. Scale bars = 20 μm.
Quantification of panel (F). Error bars indicate SEM.
DNA content analysis of Maf‐DKO macrophages infected with shRNA vectors against SIRT1 or LacZ. Each condition was done in duplicate; the results shown are representative of two independent experiments. Table indicates the percentage of cells in indicated cell cycle phases.
Quantification of panel (H), represented as ratio between proliferating (S+G2) and resting cells (G1). Data represents the pool of two independent experiments.
Analysis of colony formation potential after SIRT1 deletion by CRISPR gRNA vector infection of Cas9 expressing alveolar macrophages. Each condition was done in duplicate. Deletion efficiency of Sirt gRNA_1 and sirt gRNA_2 was evaluated by TIDE analysis (Appendix Fig S1) and corresponds to 60.9 and 44.7%, respectively. Error bars indicate SEM.

Experimental scheme of rtTA BMM differentiation and infection with tetO SIRT1 retrovirus. FACS shows purity of macrophage population as 99.8% F4/80+/CD11b+ cells.
Intracellular FACS staining of Ki67 on rtTA bone marrow‐derived macrophages (CD11+ F4/80+) infected with retrovirus expressing doxycycline‐inducible SIRT1 and constitutive GFP or empty vector (EV), 5 days post‐infection.
Quantification of panel (B). The results shown are representative of two independent experiments. Error bars indicate the standard error of the mean.

Effect of NAM on the M‐CSF‐dependent colony formation ability of Maf‐DKO macrophages. Phase contrast magnification ×10. Each condition was done in duplicate; the results shown are representative of five independent experiments. NAM was added at day 0 and colonies were counted at day 14. Scale bars = 50 μm.
Quantification of panel (A). Data represent the pool of five independent experiments.
BrdU incorporation analysis on Maf‐DKO macrophages treated or not with 10 mM NAM for 48 h. Each condition was done in duplicate; the results shown are representative of two independent experiments.
Quantification of panel (C). Data represent the pool of two independent experiments.
Immunostaining for Ki67 antigen (red) on Maf‐DKO macrophages treated or not with 10 mM NAM for 48 h. DAPI was used to stain DNA. Each condition was done in duplicate; the results shown are representative of two independent experiments. Scale bars = 20 μm.
Quantification of panel (E). At least 3,000 cells were counted per condition.
DNA content analysis of Maf‐DKO macrophages treated or not with 10 mM NAM for 48 h. Each condition was done in duplicate; the results shown are representative of two independent experiments.
Quantification of panel (G), represented as ratio between proliferating (S+G2) and resting cells (G1). Data represent the pool of two independent experiments.
Table indicates the percentage of cells in indicated cell cycle phases shown in panel (G). Error ranges indicate SEM.
Effect of different compounds NAM, the NAD+ precursor nicotinamide mononucleotide (NMN), Olaparib, a PARP inhibitor, and Inauhzin, a SIRT1‐specific antagonist on colony formation ability of Maf‐DKO macrophages. Each condition was done in duplicate; the results shown are representative of two independent experiments.
Enhanced histone H3 (Lys9) acetylation after SIRT1 inhibition with NAM. Total cell extracts were prepared from untreated or SIRT1 inhibitor (10 mM NAM, 20 h) treated Maf‐DKO macrophages. Protein blots were incubated with anti‐histone H3 (H3) or anti‐histone H3 K9acetyl (H3K9ac), as indicated. Detection of histone H3 served as controls.

Immunostaining for activated Caspase‐3 (green) on Maf‐DKO macrophages treated with NAM for 12, 24 or 48 h. Apoptotic beta islets (β) from streptozotocin‐treated mice were used as positive control. DAPI was used to stain DNA. Each condition was done in duplicate; the results shown are representative of two independent experiments. Scale bars = 20 μm.
Quantification of panel (A). At least 3,000 cells were counted per condition.
DNA content analysis of Maf‐DKO cells pre‐treated with 10 mM NAM (during 48 h) and restimulated with NAM‐free medium. Each condition was done in duplicate; the results shown are representative of three independent experiments.
Quantification of panel (C) represented as ratio between proliferating (S+G2) and resting cells (G1). Data represent the pool of three independent experiments.
Percentage of cells in indicated cell cycle phases of samples shown in panel (D). Error ranges indicate SEM.
Effect of NAM pre‐treatment (48 h) and removal on colony formation ability of Maf‐DKO macrophages. Phase contrast magnification ×10. Each condition was done in duplicate; the results shown are representative of three independent experiments.
Quantification of panel (F). Data represent the pool of three independent experiments.

Intracellular FACS staining of Ki67 on peritoneal macrophages from mice 48 h after I.P. NAM (10 mM) or control injections showing TIM4+ and TIM4− resident macrophages.
Here is the representation of two‐pooled experiments. Quantification of indicated cycling peritoneal macrophage populations expressed as a fraction of Ki67+ cells in control mice (100%). Data were pooled from two independent experiments. Percentage of Ki67+ in the respective macrophage populations are shown in Appendix Fig S2.
EdU incorporation analysis of M‐CSF stimulated MHCII+ CD11b+ F4/80+ peritoneal macrophages 48 h after NAM or control i.p. injections. Representative samples for each condition are shown (n = 2).
Quantification of panel (C). Data shown are pooled from two independent experiments using 4–6 mice per group and experiment.
Intracellular FACS staining of Ki67 of alveolar macrophages from mice 48 h after 20 mM NAM or control i.p. injections. Representative samples for each condition are shown (n = 2).
Quantification of percentage of Ki67+ cells of alveolar macrophages shown in panel (E).
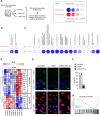
Experimental scheme of microarray Maf‐DKO macrophage sample generation and legend for BubbleGum analysis.
BubbleGUM analysis on microarray samples (Maf‐DKO macrophages treated or not with NAM for 1 or 10 h) using REACTOME mitosis gene sets.
BubbleGUM analysis on microarray samples (Maf‐DKO macrophages treated or not with NAM for 1 or 10 h) using REACTOME cell cycle phase gene sets.
Heat‐map showing the scaled expression levels (z‐score) of cyclin and cell cycle regulator related genes that were identified as coherently down‐regulated (blue) or up‐regulated (red) upon the administration of NAM (10 h).
Immunostaining for transcriptionally active phosphorylated E2F1 (p‐E2F, green) and inactive non‐phosphorylated E2F (E2F, red). 10 mM NAM was added for 48 h. DAPI was used for DNA staining. Scale bar, 20 μm. The results shown are representative of three independent experiments.
Quantification of p‐E2F1 positive cells shown as percentage of total DAPI positive cells (panel E, top). Data shown are pooled from three independent experiments. Error bars indicate SEM.
GSEA analysis of untreated versus 10‐h NAM‐treated Maf‐DKO macrophages with HALLMARK E2F transcription factor target gene set.


BubbleGUM analysis on microarray samples of Maf‐DKO macrophages treated or not with NAM for 1 h or 10 h using embryonic and tissue stem cells gene sets (Bhattacharya et al, 2004; Wong et al, 2008).
Heat‐map showing the scaled expression levels (z‐score) of self‐renewal core genes (Soucie et al, 2016) in Maf‐DKO macrophages treated or not with NAM for 1 or 10 h.
Quantitative PCR for the expression of c‐Myc, Klf2 and Klf4 in Maf‐DKO macrophages treated for the indicated times with 10 mM NAM. Average values of three independent experiments normalized to HPRT. Error bars indicate the standard error of the mean. Each condition was done in duplicate.
BubbleGUM analysis on microarray samples of Maf‐DKO macrophages treated or not with NAM for 1 or 10 h using Myc gene set data from hallmark, canonical pathways and chemical and genetic perturbations collections of the Broad Institute MSig database.

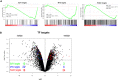
GSEA analysis of untreated versus 10‐h NAM‐treated Maf‐DKO macrophages with E2F1, FoxO1 and Myc transcription factor target gene sets. V$FOXO1_01; V$FOXO1_02; V$E2F_01; V$MYC_Q2 gene sets used.
Volcano plot analysis on microarray samples (Maf‐DKO cells treated or not with NAM for 10 h) highlighting E2F1, Fox1 and Myc target genes by GSEA (respectively, green, red and blue dots).
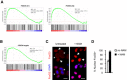
GSEA related to FOXO 4 transcriptional activity. V$FOXO4_01; V$FOXO4_02 gene sets used.
GSEA related to FoxO3 transcriptional activity (10 mM NAM 10 h).
Immunostaining for FoxO3 (red) in Maf‐DKO macrophages without or after 6 h of 10 mM NAM treatment. DAPI (blue) was used for DNA staining. Asterisks and arrows indicate cytoplasmic and nuclear localization of FoxO3, respectively. The results shown are representative of two independent experiments. Scale bars = 20 μm.
Quantification of panel (B). Averages and standard error of the mean of two independent experiments are indicated.
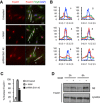
Immunofluorescence staining of FoxO1 for nuclear and cytoplasmic localization (red) in untreated Maf‐DKO macrophages or after 6‐h treatment with 10 mM NAM or after infection with shRNA SIRT1 #2 vector. FcγR (green) immunofluorescence and DAPI (blue) staining was used to define cellular and nuclear perimeters, respectively. The results shown are representative of three independent experiments. Scale bars = 20 μm.
Line profile analysis with ImageJ software of FoxO1 localization within the cells indicated with dashed line in (A). X‐ and y‐axes represent the fluorescence intensity and the position along the line used for the analyses, respectively (unit = pixels).
Quantification of nuclear FoxO1 positive cells shown as percentage of total DAPI positive cells. Data shown are pooled from three independent experiments. Error bars indicate the standard error of the mean.
FoxO1 nuclear localization after SIRT1 inhibition with NAM. Nuclear extracts or total cell extracts were prepared from untreated or SIRT1 inhibitor (10 mM NAM, 20 h) treated Maf‐DKO macrophages. Protein blots were incubated with anti‐FoxO1 as indicated. Separation of total lysates probed with anti‐FoxO1 served as controls.
References
-
- Accili D, Arden KC (2004) FoxOs at the crossroads of cellular metabolism, differentiation, and transformation. Cell 117: 421–426 - PubMed
-
- Ajami B, Bennett JL, Krieger C, Tetzlaff W, Rossi FM (2007) Local self‐renewal can sustain CNS microglia maintenance and function throughout adult life. Nat Neurosci 10: 1538–1543 - PubMed
-
- Amit I, Garber M, Chevrier N, Leite AP, Donner Y, Eisenhaure T, Guttman M, Grenier JK, Li W, Zuk O, Schubert LA, Birditt B, Shay T, Goren A, Zhang X, Smith Z, Deering R, McDonald RC, Cabili M, Bernstein BE et al (2009) Unbiased reconstruction of a mammalian transcriptional network mediating pathogen responses. Science 326: 257–263 - PMC - PubMed
-
- Avalos JL, Bever KM, Wolberger C (2005) Mechanism of sirtuin inhibition by nicotinamide: altering the NAD(+) cosubstrate specificity of a Sir2 enzyme. Mol Cell 17: 855–868 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

