Autophagy Deficiency Compromises Alternative Pathways of Respiration following Energy Deprivation in Arabidopsis thaliana
- PMID: 28710132
- PMCID: PMC5580740
- DOI: 10.1104/pp.16.01576
Autophagy Deficiency Compromises Alternative Pathways of Respiration following Energy Deprivation in Arabidopsis thaliana
Abstract
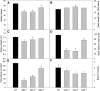
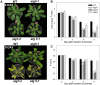
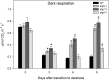
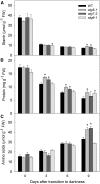
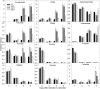
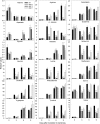
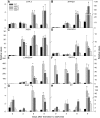
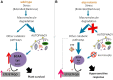
Under heterotrophic conditions, carbohydrate oxidation inside the mitochondrion is the primary energy source for cellular metabolism. However, during energy-limited conditions, alternative substrates are required to support respiration. Amino acid oxidation in plant cells plays a key role in this by generating electrons that can be transferred to the mitochondrial electron transport chain via the electron transfer flavoprotein/ubiquinone oxidoreductase system. Autophagy, a catabolic mechanism for macromolecule and protein recycling, allows the maintenance of amino acid pools and nutrient remobilization. Although the association between autophagy and alternative respiratory substrates has been suggested, the extent to which autophagy and primary metabolism interact to support plant respiration remains unclear. To investigate the metabolic importance of autophagy during development and under extended darkness, Arabidopsis (Arabidopsis thaliana) mutants with disruption of autophagy (atg mutants) were used. Under normal growth conditions, atg mutants showed lower growth and seed production with no impact on photosynthesis. Following extended darkness, atg mutants were characterized by signatures of early senescence, including decreased chlorophyll content and maximum photochemical efficiency of photosystem II coupled with increases in dark respiration. Transcript levels of genes involved in alternative pathways of respiration and amino acid catabolism were up-regulated in atg mutants. The metabolite profiles of dark-treated leaves revealed an extensive metabolic reprogramming in which increases in amino acid levels were partially compromised in atg mutants. Although an enhanced respiration in atg mutants was observed during extended darkness, autophagy deficiency compromises protein degradation and the generation of amino acids used as alternative substrates to the respiration.
© 2017 American Society of Plant Biologists. All Rights Reserved.
Figures
References
-
- Araújo WL, Ishizaki K, Nunes-Nesi A, Larson TR, Tohge T, Krahnert I, Witt S, Obata T, Schauer N, Graham IA, et al. (2010) Identification of the 2-hydroxyglutarate and isovaleryl-CoA dehydrogenases as alternative electron donors linking lysine catabolism to the electron transport chain of Arabidopsis mitochondria. Plant Cell 22: 1549–1563 - PMC - PubMed
-
- Araújo WL, Tohge T, Ishizaki K, Leaver CJ, Fernie AR (2011) Protein degradation: an alternative respiratory substrate for stressed plants. Trends Plant Sci 16: 489–498 - PubMed
-
- Avila-Ospina L, Moison M, Yoshimoto K, Masclaux-Daubresse C (2014) Autophagy, plant senescence, and nutrient recycling. J Exp Bot 65: 3799–3811 - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

