Cathelicidins Inhibit Escherichia coli-Induced TLR2 and TLR4 Activation in a Viability-Dependent Manner
- PMID: 28710255
- PMCID: PMC5544931
- DOI: 10.4049/jimmunol.1602164
Cathelicidins Inhibit Escherichia coli-Induced TLR2 and TLR4 Activation in a Viability-Dependent Manner
Abstract
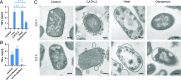
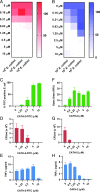
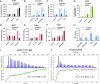
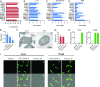
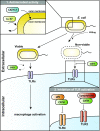
Activation of the immune system needs to be tightly regulated to provide protection against infections and, at the same time, to prevent excessive inflammation to limit collateral damage to the host. This tight regulation includes regulating the activation of TLRs, which are key players in the recognition of invading microbes. A group of short cationic antimicrobial peptides, called cathelicidins, have previously been shown to modulate TLR activation by synthetic or purified TLR ligands and may play an important role in the regulation of inflammation during infections. However, little is known about how these cathelicidins affect TLR activation in the context of complete and viable bacteria. In this article, we show that chicken cathelicidin-2 kills Escherichia coli in an immunogenically silent fashion. Our results show that chicken cathelicidin-2 kills E. coli by permeabilizing the bacterial inner membrane and subsequently binds the outer membrane-derived lipoproteins and LPS to inhibit TLR2 and TLR4 activation, respectively. In addition, other cathelicidins, including human, mouse, pig, and dog cathelicidins, which lack antimicrobial activity under cell culture conditions, only inhibit macrophage activation by nonviable E. coli In total, this study shows that cathelicidins do not affect immune activation by viable bacteria and only inhibit inflammation when bacterial viability is lost. Therefore, cathelicidins provide a novel mechanism by which the immune system can discriminate between viable and nonviable Gram-negative bacteria to tune the immune response, thereby limiting collateral damage to the host and the risk for sepsis.
Copyright © 2017 by The American Association of Immunologists, Inc.
Figures

References
-
- Gewirtz A. T., Navas T. A., Lyons S., Godowski P. J., Madara J. L. 2001. Cutting edge: bacterial flagellin activates basolaterally expressed TLR5 to induce epithelial proinflammatory gene expression. J. Immunol. 167: 1882–1885. - PubMed
-
- Soehnlein O., Lindbom L. 2010. Phagocyte partnership during the onset and resolution of inflammation. Nat. Rev. Immunol. 10: 427–439. - PubMed
-
- Liew F. Y., Xu E. K., Brint, O’Neill L. A. J. 2005. Negative regulation of toll-like receptor-mediated immune responses. Nat. Rev. Immunol. 5: 446–458. - PubMed
-
- Nizet V., Ohtake T., Lauth X., Trowbridge J., Rudisill J., Dorschner R. A., Pestonjamasp V., Piraino J., Huttner K., Gallo R. L. 2001. Innate antimicrobial peptide protects the skin from invasive bacterial infection. Nature 414: 454–457. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

