Unsupervised Extraction of Stable Expression Signatures from Public Compendia with an Ensemble of Neural Networks
- PMID: 28711280
- PMCID: PMC5532071
- DOI: 10.1016/j.cels.2017.06.003
Unsupervised Extraction of Stable Expression Signatures from Public Compendia with an Ensemble of Neural Networks
Abstract
Cross-experiment comparisons in public data compendia are challenged by unmatched conditions and technical noise. The ADAGE method, which performs unsupervised integration with denoising autoencoder neural networks, can identify biological patterns, but because ADAGE models, like many neural networks, are over-parameterized, different ADAGE models perform equally well. To enhance model robustness and better build signatures consistent with biological pathways, we developed an ensemble ADAGE (eADAGE) that integrated stable signatures across models. We applied eADAGE to a compendium of Pseudomonas aeruginosa gene expression profiling experiments performed in 78 media. eADAGE revealed a phosphate starvation response controlled by PhoB in media with moderate phosphate and predicted that a second stimulus provided by the sensor kinase, KinB, is required for this PhoB activation. We validated this relationship using both targeted and unbiased genetic approaches. eADAGE, which captures stable biological patterns, enables cross-experiment comparisons that can highlight measured but undiscovered relationships.
Keywords: Pseudomonas aeruginosa; crosstalk; denoising autoencoders; ensemble modeling; gene expression; neural networks; phosphate starvation.
Copyright © 2017 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures
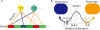



References
-
- Beaulieu-Jones BK, Greene CS Pooled Resource Open-Access ALS Clinical Trials Consortium. Semi-supervised learning of the electronic health record for phenotype stratification. J. Biomed. Inform. 2016;64:168–178. - PubMed
-
- Bengio Y, Courville A, Vincent P. Representation Learning: A Review and New Perspectives. TPAMI. 2013;35:1798–1828. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

