Diversity and distribution of nuclease bacteriocins in bacterial genomes revealed using Hidden Markov Models
- PMID: 28715501
- PMCID: PMC5536347
- DOI: 10.1371/journal.pcbi.1005652
Diversity and distribution of nuclease bacteriocins in bacterial genomes revealed using Hidden Markov Models
Abstract
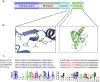
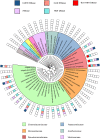
Bacteria exploit an arsenal of antimicrobial peptides and proteins to compete with each other. Three main competition systems have been described: type six secretion systems (T6SS); contact dependent inhibition (CDI); and bacteriocins. Unlike T6SS and CDI systems, bacteriocins do not require contact between bacteria but are diffusible toxins released into the environment. Identified almost a century ago, our understanding of bacteriocin distribution and prevalence in bacterial populations remains poor. In the case of protein bacteriocins, this is because of high levels of sequence diversity and difficulties in distinguishing their killing domains from those of other competition systems. Here, we develop a robust bioinformatics pipeline exploiting Hidden Markov Models for the identification of nuclease bacteriocins (NBs) in bacteria of which, to-date, only a handful are known. NBs are large (>60 kDa) toxins that target nucleic acids (DNA, tRNA or rRNA) in the cytoplasm of susceptible bacteria, usually closely related to the producing organism. We identified >3000 NB genes located on plasmids or on the chromosome from 53 bacterial species distributed across different ecological niches, including human, animals, plants, and the environment. A newly identified NB predicted to be specific for Pseudomonas aeruginosa (pyocin Sn) was produced and shown to kill P. aeruginosa thereby validating our pipeline. Intriguingly, while the genes encoding the machinery needed for NB translocation across the cell envelope are widespread in Gram-negative bacteria, NBs are found exclusively in γ-proteobacteria. Similarity network analysis demonstrated that NBs fall into eight groups each with a distinct arrangement of protein domains involved in import. The only structural feature conserved across all groups was a sequence motif critical for cell-killing that is generally not found in bacteriocins targeting the periplasm, implying a specific role in translocating the nuclease to the cytoplasm. Finally, we demonstrate a significant association between nuclease colicins, NBs specific for Escherichia coli, and virulence factors, suggesting NBs play a role in infection processes, most likely by enabling pathogens to outcompete commensal bacteria.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Gratia A. Sur un remarquable exemple d'antagonisme entre deux souches de colibacille. CR SocBiol (Paris). 1925;(93):1040–1.
-
- Kerr B, Riley MA, Feldman MW, Bohannan BJ. Local dispersal promotes biodiversity in a real-life game of rock-paper-scissors. Nature. 2002;418(6894):171–4. doi: 10.1038/nature00823 . - DOI - PubMed
-
- Majeed H, Gillor O, Kerr B, Riley Ma. Competitive interactions in Escherichia coli populations: the role of bacteriocins. The ISME journal. 2011;5:71–81. doi: 10.1038/ismej.2010.90 . - DOI - PMC - PubMed
-
- Majeed H, Lampert A, Ghazaryan L, Gillor O. The weak shall inherit: bacteriocin-mediated interactions in bacterial populations. PloS one. 2013;8:e63837 doi: 10.1371/journal.pone.0063837 . - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

