The Evolving Genomic Landscape of Barrett's Esophagus and Esophageal Adenocarcinoma
- PMID: 28716721
- PMCID: PMC6025803
- DOI: 10.1053/j.gastro.2017.07.007
The Evolving Genomic Landscape of Barrett's Esophagus and Esophageal Adenocarcinoma
Abstract
We have recently gained unprecedented insight into genetic factors that determine risk for Barrett's esophagus (BE) and progression to esophageal adenocarcinoma (EA). Next-generation sequencing technologies have allowed us to identify somatic mutations that initiate BE and track genetic changes during development of tumors and invasive cancer. These technologies led to identification of mechanisms of tumorigenesis that challenge the current multistep model of progression to EA. Newer, cost-effective technologies create opportunities to rapidly translate the analysis of DNA into tools that can identify patients with BE at high risk for cancer, detect dysplastic lesions more reliably, and uncover mechanisms of carcinogenesis.
Keywords: Chromothripsis; Cytosponge; Esophagus; Genome-wide Association Study; Mutational Signature.
Copyright © 2017 AGA Institute. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
RCF is named on patents pertaining to the Cytopsonge and associated assays that have been licensed by the Medical Research Council to Covidien (now Medtronic). All the authors declare no other competing interests.
Figures
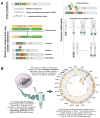




References
-
- National Cancer Institute. Cancer of the Esophagus - SEER Stat Fact Sheets. Cancer Stat. 2015 Available at: http://seer.cancer.gov/statfacts/html/esoph.html.
-
- Cancer Resarch UK. Oesophageal cancer incidence statistics. Available at: cancerresearchuk.org/cancer-info/cancerstats/
-
- Hirst J, Smithers BM, Gotley DC, et al. Defining cure for esophageal cancer: analysis of actual 5-year survivors following esophagectomy. Ann Surg Oncol. 2011;18:1766–74. - PubMed
-
- van Hagen P, Hulshof MCCM, van Lanschot JJB, et al. Preoperative Chemoradiotherapy for Esophageal or Junctional Cancer. N Engl J Med. 2012;366:2074–2084. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

