Direct interaction of the Golgi V-ATPase a-subunit isoform with PI(4)P drives localization of Golgi V-ATPases in yeast
- PMID: 28720663
- PMCID: PMC5597324
- DOI: 10.1091/mbc.E17-05-0316
Direct interaction of the Golgi V-ATPase a-subunit isoform with PI(4)P drives localization of Golgi V-ATPases in yeast
Abstract
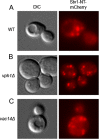
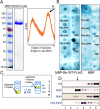
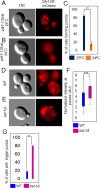
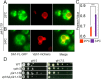
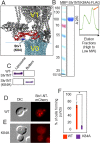
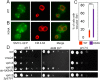
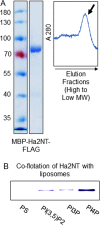
Luminal pH and phosphoinositide content are fundamental features of organelle identity. Vacuolar H+-ATPases (V-ATPases) drive organelle acidification in all eukaryotes, and membrane-bound a-subunit isoforms of the V-ATPase are implicated in organelle-specific targeting and regulation. Earlier work demonstrated that the endolysosomal lipid PI(3,5)P2 activates V-ATPases containing the vacuolar a-subunit isoform in Saccharomyces cerevisiae Here we demonstrate that PI(4)P, the predominant Golgi phosphatidylinositol (PI) species, directly interacts with the cytosolic amino terminal (NT) domain of the yeast Golgi V-ATPase a-isoform Stv1. Lysine-84 of Stv1NT is essential for interaction with PI(4)P in vitro and in vivo, and interaction with PI(4)P is required for efficient localization of Stv1-containing V-ATPases. The cytosolic NT domain of the human V-ATPase a2 isoform specifically interacts with PI(4)P in vitro, consistent with its Golgi localization and function. We propose that NT domains of Vo a-subunit isoforms interact specifically with PI lipids in their organelles of residence. These interactions can transmit organelle-specific targeting or regulation information to V-ATPases.
© 2017 Banerjee and Kane. This article is distributed by The American Society for Cell Biology under license from the author(s). Two months after publication it is available to the public under an Attribution–Noncommercial–Share Alike 3.0 Unported Creative Commons License (http://creativecommons.org/licenses/by-nc-sa/3.0).
Figures
Similar articles
-
Human V-ATPase a-subunit isoforms bind specifically to distinct phosphoinositide phospholipids.J Biol Chem. 2023 Dec;299(12):105473. doi: 10.1016/j.jbc.2023.105473. Epub 2023 Nov 17. J Biol Chem. 2023. PMID: 37979916 Free PMC article.
-
Interaction of the late endo-lysosomal lipid PI(3,5)P2 with the Vph1 isoform of yeast V-ATPase increases its activity and cellular stress tolerance.J Biol Chem. 2019 Jun 7;294(23):9161-9171. doi: 10.1074/jbc.RA119.008552. Epub 2019 Apr 25. J Biol Chem. 2019. PMID: 31023825 Free PMC article.
-
The RAVE complex is an isoform-specific V-ATPase assembly factor in yeast.Mol Biol Cell. 2014 Feb;25(3):356-67. doi: 10.1091/mbc.E13-05-0231. Epub 2013 Dec 4. Mol Biol Cell. 2014. PMID: 24307682 Free PMC article.
-
Regulation of V-ATPase Activity and Organelle pH by Phosphatidylinositol Phosphate Lipids.Front Cell Dev Biol. 2020 Jun 23;8:510. doi: 10.3389/fcell.2020.00510. eCollection 2020. Front Cell Dev Biol. 2020. PMID: 32656214 Free PMC article. Review.
-
The cytosolic N-terminal domain of V-ATPase a-subunits is a regulatory hub targeted by multiple signals.Front Mol Biosci. 2023 Jun 16;10:1168680. doi: 10.3389/fmolb.2023.1168680. eCollection 2023. Front Mol Biosci. 2023. PMID: 37398550 Free PMC article. Review.
Cited by
-
The Role of Sch9 and the V-ATPase in the Adaptation Response to Acetic Acid and the Consequences for Growth and Chronological Lifespan.Microorganisms. 2021 Sep 3;9(9):1871. doi: 10.3390/microorganisms9091871. Microorganisms. 2021. PMID: 34576766 Free PMC article.
-
Non-autophagic Golgi-LC3 lipidation facilitates TFE3 stress response against Golgi dysfunction.EMBO J. 2024 Nov;43(21):5085-5113. doi: 10.1038/s44318-024-00233-y. Epub 2024 Sep 16. EMBO J. 2024. PMID: 39284911 Free PMC article.
-
Chimeric a-subunit isoforms generate functional yeast V-ATPases with altered regulatory properties in vitro and in vivo.Mol Biol Cell. 2023 Mar 1;34(3):ar14. doi: 10.1091/mbc.E22-07-0265. Epub 2023 Jan 4. Mol Biol Cell. 2023. PMID: 36598799 Free PMC article.
-
Yeast TLDc domain proteins regulate assembly state and subcellular localization of the V-ATPase.EMBO J. 2024 May;43(9):1870-1897. doi: 10.1038/s44318-024-00097-2. Epub 2024 Apr 8. EMBO J. 2024. PMID: 38589611 Free PMC article.
-
Vacuolar-type ATPase: A proton pump to lysosomal trafficking.Proc Jpn Acad Ser B Phys Biol Sci. 2019;95(6):261-277. doi: 10.2183/pjab.95.018. Proc Jpn Acad Ser B Phys Biol Sci. 2019. PMID: 31189779 Free PMC article. Review.
References
-
- Audhya A, Emr SD. Stt4 PI 4-kinase localizes to the plasma membrane and functions in the Pkc1-mediated MAP kinase cascade. Dev Cell. 2002;2:593–605. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

