Repeat-Induced Point Mutation and Other Genome Defense Mechanisms in Fungi
- PMID: 28721856
- PMCID: PMC5607778
- DOI: 10.1128/microbiolspec.FUNK-0042-2017
Repeat-Induced Point Mutation and Other Genome Defense Mechanisms in Fungi
Abstract
Transposable elements have colonized the genomes of nearly all organisms, including fungi. Although transposable elements may sometimes provide beneficial functions to their hosts their overall impact is considered deleterious. As a result, the activity of transposable elements needs to be counterbalanced by the host genome defenses. In fungi, the primary genome defense mechanisms include repeat-induced point mutation (RIP) and methylation induced premeiotically, meiotic silencing by unpaired DNA, sex-induced silencing, cosuppression (also known as somatic quelling), and cotranscriptional RNA surveillance. Recent studies of the filamentous fungus Neurospora crassa have shown that the process of repeat recognition for RIP apparently involves interactions between coaligned double-stranded segments of chromosomal DNA. These studies have also shown that RIP can be mediated by the conserved pathway that establishes transcriptional (heterochromatic) silencing of repetitive DNA. In light of these new findings, RIP emerges as a specialized case of the general phenomenon of heterochromatic silencing of repetitive DNA.
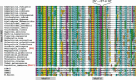
Figures

Similar articles
-
RIP: the evolutionary cost of genome defense.Trends Genet. 2004 Sep;20(9):417-23. doi: 10.1016/j.tig.2004.07.007. Trends Genet. 2004. PMID: 15313550 Review.
-
Recombination-independent recognition of DNA homology for repeat-induced point mutation.Curr Genet. 2017 Jun;63(3):389-400. doi: 10.1007/s00294-016-0649-4. Epub 2016 Sep 14. Curr Genet. 2017. PMID: 27628707 Free PMC article. Review.
-
Repeat-induced point (RIP) mutation in the industrial workhorse fungus Trichoderma reesei.Appl Microbiol Biotechnol. 2018 Feb;102(4):1567-1574. doi: 10.1007/s00253-017-8731-5. Epub 2018 Jan 8. Appl Microbiol Biotechnol. 2018. PMID: 29308529 Review.
-
The evolution of transposon repeat-induced point mutation in the genome of Colletotrichum cereale: reconciling sex, recombination and homoplasy in an ''asexual" pathogen.Fungal Genet Biol. 2008 Mar;45(3):190-206. doi: 10.1016/j.fgb.2007.08.004. Epub 2007 Aug 29. Fungal Genet Biol. 2008. PMID: 17962053
-
Gene inactivation triggered by recognition between DNA repeats.Experientia. 1994 Mar 15;50(3):307-17. doi: 10.1007/BF01924014. Experientia. 1994. PMID: 8143804 Review.
Cited by
-
Encyclopaedia of eukaryotic DNA methylation: from patterns to mechanisms and functions.Biochem Soc Trans. 2022 Jun 30;50(3):1179-1190. doi: 10.1042/BST20210725. Biochem Soc Trans. 2022. PMID: 35521905 Free PMC article.
-
The model diatom Phaeodactylum tricornutum provides insights into the diversity and function of microeukaryotic DNA methyltransferases.Commun Biol. 2023 Mar 9;6(1):253. doi: 10.1038/s42003-023-04629-0. Commun Biol. 2023. PMID: 36894681 Free PMC article.
-
Population genomics of transposable element activation in the highly repressive genome of an agricultural pathogen.Microb Genom. 2021 Aug;7(8):000540. doi: 10.1099/mgen.0.000540. Microb Genom. 2021. PMID: 34424154 Free PMC article.
-
A DEAD-box RNA helicase mediates meiotic silencing by unpaired DNA.G3 (Bethesda). 2023 Aug 9;13(8):jkad083. doi: 10.1093/g3journal/jkad083. G3 (Bethesda). 2023. PMID: 37052947 Free PMC article.
-
Genome-Wide Analyses of Repeat-Induced Point Mutations in the Ascomycota.Front Microbiol. 2021 Feb 1;11:622368. doi: 10.3389/fmicb.2020.622368. eCollection 2020. Front Microbiol. 2021. PMID: 33597932 Free PMC article.
References
-
- Castanera R, López-Varas L, Borgognone A, LaButti K, Lapidus A, Schmutz J, Grimwood J, Pérez G, Pisabarro AG, Grigoriev IV, Stajich JE, Ramírez L. 2016. Transposable elements versus the fungal genome: impact on whole-genome architecture and transcriptional profiles. PLoS Genet 12:e1006108. 10.1371/journal.pgen.1006108 - DOI - PMC - PubMed
-
- Ohm RA, Feau N, Henrissat B, Schoch CL, Horwitz BA, Barry KW, Condon BJ, Copeland AC, Dhillon B, Glaser F, Hesse CN, Kosti I, LaButti K, Lindquist EA, Lucas S, Salamov AA, Bradshaw RE, Ciuffetti L, Hamelin RC, Kema GH, Lawrence C, Scott JA, Spatafora JW, Turgeon BG, de Wit PJ, Zhong S, Goodwin SB, Grigoriev IV. 2012. Diverse lifestyles and strategies of plant pathogenesis encoded in the genomes of eighteen Dothideomycetes fungi. PLoS Pathog 8:e1003037. 10.1371/journal.ppat.1003037 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

