The Metal Drives the Chemistry: Dual Functions of Acireductone Dioxygenase
- PMID: 28731690
- PMCID: PMC5604235
- DOI: 10.1021/acs.chemrev.7b00117
The Metal Drives the Chemistry: Dual Functions of Acireductone Dioxygenase
Abstract
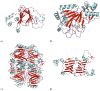
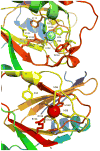
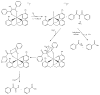
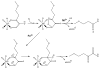
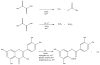
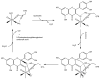
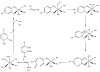
Acireductone dioxygenase (ARD) from the methionine salvage pathway (MSP) is a unique enzyme that exhibits dual chemistry determined solely by the identity of the divalent transition-metal ion (Fe2+ or Ni2+) in the active site. The Fe2+-containing isozyme catalyzes the on-pathway reaction using substrates 1,2-dihydroxy-3-keto-5-methylthiopent-1-ene (acireductone) and dioxygen to generate formate and the ketoacid precursor of methionine, 2-keto-4-methylthiobutyrate, whereas the Ni2+-containing isozyme catalyzes an off-pathway shunt with the same substrates, generating methylthiopropionate, carbon monoxide, and formate. The dual chemistry of ARD was originally discovered in the bacterium Klebsiella oxytoca, but it has recently been shown that mammalian ARD enzymes (mouse and human) are also capable of catalyzing metal-dependent dual chemistry in vitro. This is particularly interesting, since carbon monoxide, one of the products of off-pathway reaction, has been identified as an antiapoptotic molecule in mammals. In addition, several biochemical and genetic studies have indicated an inhibitory role of human ARD in cancer. This comprehensive review describes the biochemical and structural characterization of the ARD family, the proposed experimental and theoretical approaches to establishing mechanisms for the dual chemistry, insights into the mechanism based on comparison with structurally and functionally similar enzymes, and the applications of this research to the field of artificial metalloenzymes and synthetic biology.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

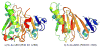
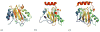
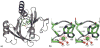
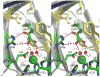

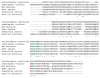
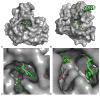

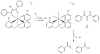
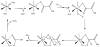
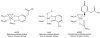
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

