The T-cell leukemia-associated ribosomal RPL10 R98S mutation enhances JAK-STAT signaling
- PMID: 28744013
- PMCID: PMC5669462
- DOI: 10.1038/leu.2017.225
The T-cell leukemia-associated ribosomal RPL10 R98S mutation enhances JAK-STAT signaling
Abstract
Several somatic ribosome defects have recently been discovered in cancer, yet their oncogenic mechanisms remain poorly understood. Here we investigated the pathogenic role of the recurrent R98S mutation in ribosomal protein L10 (RPL10 R98S) found in T-cell acute lymphoblastic leukemia (T-ALL). The JAK-STAT signaling pathway is a critical controller of cellular proliferation and survival. A proteome screen revealed overexpression of several Jak-Stat signaling proteins in engineered RPL10 R98S mouse lymphoid cells, which we confirmed in hematopoietic cells from transgenic Rpl10 R98S mice and T-ALL xenograft samples. RPL10 R98S expressing cells displayed JAK-STAT pathway hyper-activation upon cytokine stimulation, as well as increased sensitivity to clinically used JAK-STAT inhibitors like pimozide. A mutually exclusive mutation pattern between RPL10 R98S and JAK-STAT mutations in T-ALL patients further suggests that RPL10 R98S functionally mimics JAK-STAT activation. Mechanistically, besides transcriptional changes, RPL10 R98S caused reduction of apparent programmed ribosomal frameshifting at several ribosomal frameshift signals in mouse and human Jak-Stat genes, as well as decreased Jak1 degradation. Of further medical interest, RPL10 R98S cells showed reduced proteasome activity and enhanced sensitivity to clinical proteasome inhibitors. Collectively, we describe modulation of the JAK-STAT cascade as a novel cancer-promoting activity of a ribosomal mutation, and expand the relevance of this cascade in leukemia.
Conflict of interest statement
The authors disclose no potential conflicts of interest.
Figures



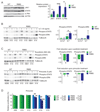


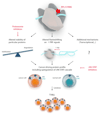
References
-
- Pui C-H, Robison LL, Look AT. Acute lymphoblastic leukaemia. The Lancet. 2008;371:1030–1043. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

