Redundant and diverse intranodal pacemakers and conduction pathways protect the human sinoatrial node from failure
- PMID: 28747516
- PMCID: PMC5775890
- DOI: 10.1126/scitranslmed.aam5607
Redundant and diverse intranodal pacemakers and conduction pathways protect the human sinoatrial node from failure
Abstract
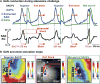
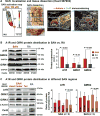
The human sinoatrial node (SAN) efficiently maintains heart rhythm even under adverse conditions. However, the specific mechanisms involved in the human SAN's ability to prevent rhythm failure, also referred to as its robustness, are unknown. Challenges exist because the three-dimensional (3D) intramural structure of the human SAN differs from well-studied animal models, and clinical electrode recordings are limited to only surface atrial activation. Hence, to innovate the translational study of human SAN structural and functional robustness, we integrated intramural optical mapping, 3D histology reconstruction, and molecular mapping of the ex vivo human heart. When challenged with adenosine or atrial pacing, redundant intranodal pacemakers within the human SAN maintained automaticity and delivered electrical impulses to the atria through sinoatrial conduction pathways (SACPs), thereby ensuring a fail-safe mechanism for robust maintenance of sinus rhythm. During adenosine perturbation, the primary central SAN pacemaker was suppressed, whereas previously inactive superior or inferior intranodal pacemakers took over automaticity maintenance. Sinus rhythm was also rescued by activation of another SACP when the preferential SACP was suppressed, suggesting two independent fail-safe mechanisms for automaticity and conduction. The fail-safe mechanism in response to adenosine challenge is orchestrated by heterogeneous differences in adenosine A1 receptors and downstream GIRK4 channel protein expressions across the SAN complex. Only failure of all pacemakers and/or SACPs resulted in SAN arrest or conduction block. Our results unmasked reserve mechanisms that protect the human SAN pacemaker and conduction complex from rhythm failure, which may contribute to treatment of SAN arrhythmias.
Copyright © 2017 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Conflict of interest statement
Figures


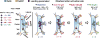

Comment in
-
Arrhythmias: Safety backups to keep the pace.Nat Rev Cardiol. 2017 Aug 16;14(9):503. doi: 10.1038/nrcardio.2017.126. Nat Rev Cardiol. 2017. PMID: 28812711 No abstract available.
References
-
- Chandler NJ, Greener ID, Tellez JO, Inada S, Musa H, Molenaar P, DiFrancesco D, Baruscotti M, Longhi R, Anderson RH, Billeter R, Sharma V, Sigg DC, Boyett MR, Dobrzynski H. Molecular architecture of the human sinus node: Insights into the function of the cardiac pacemaker. Circulation. 2009;119:1562–1575. - PubMed
-
- Fedorov VV, Glukhov AV, Chang R. Conduction barriers and pathways of the sinoatrial pacemaker complex: Their role in normal rhythm and atrial arrhythmias. Am J Physiol Heart Circ Physiol. 2012;302:H1773–H1783. - PubMed
-
- Gomes JA, Winters SL. The origins of the sinus node pacemaker complex in man: Demonstration of dominant and subsidiary foci. J Am Coll Cardiol. 1987;9:45–52. - PubMed
-
- Boineau JP, Canavan TE, Schuessler RB, Cain ME, Corr PB, Cox JL. Demonstration of a widely distributed atrial pacemaker complex in the human heart. Circulation. 1988;77:1221–1237. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

