Targeted next-generation sequencing detects novel gene-phenotype associations and expands the mutational spectrum in cardiomyopathies
- PMID: 28750076
- PMCID: PMC5531468
- DOI: 10.1371/journal.pone.0181842
Targeted next-generation sequencing detects novel gene-phenotype associations and expands the mutational spectrum in cardiomyopathies
Abstract
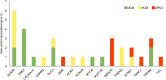
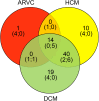
Cardiomyopathies are a heterogeneous group of primary diseases of the myocardium, including hypertrophic cardiomyopathy (HCM), dilated cardiomyopathy (DCM), and arrhythmogenic right ventricular cardiomyopathy (ARVC), with higher morbidity and mortality. These diseases are genetically diverse and associated with rare mutations in a large number of genes, many of which overlap among the phenotypes. To better investigate the genetic overlap between these three phenotypes and to identify new genotype-phenotype correlations, we designed a custom gene panel consisting of 115 genes known to be associated with cardiomyopathic phenotypes and channelopathies. A cohort of 38 unrelated patients, 16 affected by DCM, 14 by HCM and 8 by ARVC, was recruited for the study on the basis of more severe phenotypes and family history of cardiomyopathy and/or sudden death. We detected a total of 142 rare variants in 40 genes, and all patients were found to be carriers of at least one rare variant. Twenty-eight of the 142 rare variants were also predicted as potentially pathogenic variants and found in 26 patients. In 23 out of 38 patients, we found at least one novel potential gene-phenotype association. In particular, we detected three variants in OBSCN gene in ARVC patients, four variants in ANK2 gene and two variants in DLG1, TRPM4, and AKAP9 genes in DCM patients, two variants in PSEN2 gene and four variants in AKAP9 gene in HCM patients. Overall, our results confirmed that cardiomyopathic patients could carry multiple rare gene variants; in addition, our investigation of the genetic overlap among cardiomyopathies revealed new gene-phenotype associations. Furthermore, as our study confirms, data obtained using targeted next-generation sequencing could provide a remarkable contribution to the molecular diagnosis of cardiomyopathies, early identification of patients at risk for arrhythmia development, and better clinical management of cardiomyopathic patients.
Conflict of interest statement
Figures
References
-
- Elliott P, Andersson B, Arbustini E, Bilinska Z, Cecchi F, Charron P, et al. Classification of the cardiomyopathies: a position statement from the European Society Of Cardiology Working Group on Myocardial and Pericardial Diseases. Eur Heart J. 2008;29(2):270–6. doi: 10.1093/eurheartj/ehm342 - DOI - PubMed
-
- Maron BJ, Towbin JA, Thiene G, Antzelevitch C, Corrado D, Arnett D, et al. Contemporary definitions and classification of the cardiomyopathies: an American Heart Association Scientific Statement from the Council on Clinical Cardiology, Heart Failure and Transplantation Committee; Quality of Care and Outcomes Research and Functional Genomics and Translational Biology Interdisciplinary Working Groups; and Council on Epidemiology and Prevention. Circulation. 2006;113(14):1807–16. doi: 10.1161/CIRCULATIONAHA.106.174287 - DOI - PubMed
-
- Authors/Task Force m, Elliott PM, Anastasakis A, Borger MA, Borggrefe M, Cecchi F, et al. 2014 ESC Guidelines on diagnosis and management of hypertrophic cardiomyopathy: the Task Force for the Diagnosis and Management of Hypertrophic Cardiomyopathy of the European Society of Cardiology (ESC). Eur Heart J. 2014;35(39):2733–79. doi: 10.1093/eurheartj/ehu284 - DOI - PubMed
-
- Elliott P, McKenna WJ. Hypertrophic cardiomyopathy. Lancet. 2004;363(9424):1881–91. doi: 10.1016/S0140-6736(04)16358-7 - DOI - PubMed
-
- Cahill TJ, Ashrafian H, Watkins H. Genetic cardiomyopathies causing heart failure. Circ Res. 2013;113(6):660–75. doi: 10.1161/CIRCRESAHA.113.300282 - DOI - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical