Quantitative Proteomic Analysis of Optimal Cutting Temperature (OCT) Embedded Core-Needle Biopsy of Lung Cancer
- PMID: 28752479
- PMCID: PMC5693617
- DOI: 10.1007/s13361-017-1706-z
Quantitative Proteomic Analysis of Optimal Cutting Temperature (OCT) Embedded Core-Needle Biopsy of Lung Cancer
Abstract
With recent advances in understanding the genomic underpinnings and oncogenic drivers of pathogenesis in different subtypes, it is increasingly clear that proper pretreatment diagnostics are essential for the choice of appropriate treatment options for non-small cell lung cancer (NSCLC). Tumor tissue preservation in optimal cutting temperature (OCT) compound is commonly used in the surgical suite. However, proteins recovered from OCT-embedded specimens pose a challenge for LC-MS/MS experiments, due to the large amounts of polymers present in OCT. Here we present a simple workflow for whole proteome analysis of OCT-embedded NSCLC tissue samples, which involves a simple trichloroacetic acid precipitation step. Comparisons of protein recovery between frozen versus OCT-embedded tissue showed excellent consistency with more than 9200 proteins identified. Using an isobaric labeling strategy, we quantified more than 5400 proteins in tumor versus normal OCT-embedded core needle biopsy samples. Gene ontology analysis indicated that a number of proliferative as well as squamous cell carcinoma (SqCC) marker proteins were overexpressed in the tumor, consistent with the patient's pathology based diagnosis of "poorly differentiated SqCC". Among the most downregulated proteins in the tumor sample, we noted a number of proteins with potential immunomodulatory functions. Finally, interrogation of the aberrantly expressed proteins using a candidate approach and cross-referencing with publicly available databases led to the identification of potential druggable targets in DNA replication and DNA damage repair pathways. We conclude that our approach allows LC-MS/MS proteomic analyses on OCT-embedded lung cancer specimens, opening the way to bring powerful proteomics into the clinic. Graphical Abstract ᅟ.
Keywords: Biomarker; Drug target; Lung cancer; Pathology; Proteomics; Sample preparation.
Figures




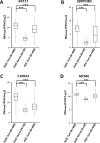
References
-
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66:7–30. - PubMed
-
- Travis WD. Pathology of lung cancer. Clin Chest Med. 2011;32:669–692. - PubMed
-
- Johnson DH, Fehrenbacher L, Novotny WF, Herbst RS, Nemunaitis JJ, Jablons DM, Langer CJ, DeVore RF, 3rd, Gaudreault J, Damico LA, Holmgren E, Kabbinavar F. Randomized phase II trial comparing bevacizumab plus carboplatin and paclitaxel with carboplatin and paclitaxel alone in previously untreated locally advanced or metastatic non-small-cell lung cancer. J Clin Oncol. 2004;22:2184–2191. - PubMed
-
- Pesch B, Kendzia B, Gustavsson P, Jockel KH, Johnen G, Pohlabeln H, Olsson A, Ahrens W, Gross IM, Bruske I, Wichmann HE, Merletti F, Richiardi L, Simonato L, Fortes C, Siemiatycki J, Parent ME, Consonni D, Landi MT, Caporaso N, Zaridze D, Cassidy A, Szeszenia-Dabrowska N, Rudnai P, Lissowska J, Stucker I, Fabianova E, Dumitru RS, Bencko V, Foretova L, Janout V, Rudin CM, Brennan P, Boffetta P, Straif K, Bruning T. Cigarette smoking and lung cancer–relative risk estimates for the major histological types from a pooled analysis of case-control studies. Int J Cancer. 2012;131:1210–1219. - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

