The Drosophila Duox maturation factor is a key component of a positive feedback loop that sustains regeneration signaling
- PMID: 28753614
- PMCID: PMC5550008
- DOI: 10.1371/journal.pgen.1006937
The Drosophila Duox maturation factor is a key component of a positive feedback loop that sustains regeneration signaling
Abstract
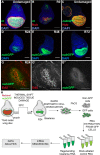
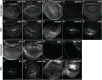
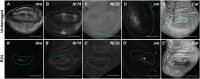
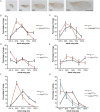
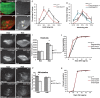
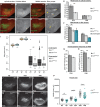
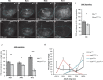
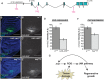
Regenerating tissue must initiate the signaling that drives regenerative growth, and sustain that signaling long enough for regeneration to complete. How these key signals are sustained is unclear. To gain a comprehensive view of the changes in gene expression that occur during regeneration, we performed whole-genome mRNAseq of actively regenerating tissue from damaged Drosophila wing imaginal discs. We used genetic tools to ablate the wing primordium to induce regeneration, and carried out transcriptional profiling of the regeneration blastema by fluorescently labeling and sorting the blastema cells, thus identifying differentially expressed genes. Importantly, by using genetic mutants of several of these differentially expressed genes we have confirmed that they have roles in regeneration. Using this approach, we show that high expression of the gene moladietz (mol), which encodes the Duox-maturation factor NIP, is required during regeneration to produce reactive oxygen species (ROS), which in turn sustain JNK signaling during regeneration. We also show that JNK signaling upregulates mol expression, thereby activating a positive feedback signal that ensures the prolonged JNK activation required for regenerative growth. Thus, by whole-genome transcriptional profiling of regenerating tissue we have identified a positive feedback loop that regulates the extent of regenerative growth.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Similar articles
-
Cap-n-Collar Promotes Tissue Regeneration by Regulating ROS and JNK Signaling in the Drosophila melanogaster Wing Imaginal Disc.Genetics. 2017 Jul;206(3):1505-1520. doi: 10.1534/genetics.116.196832. Epub 2017 May 16. Genetics. 2017. PMID: 28512185 Free PMC article.
-
ROS-Induced JNK and p38 Signaling Is Required for Unpaired Cytokine Activation during Drosophila Regeneration.PLoS Genet. 2015 Oct 23;11(10):e1005595. doi: 10.1371/journal.pgen.1005595. eCollection 2015 Oct. PLoS Genet. 2015. PMID: 26496642 Free PMC article.
-
Ask1 and Akt act synergistically to promote ROS-dependent regeneration in Drosophila.PLoS Genet. 2019 Jan 24;15(1):e1007926. doi: 10.1371/journal.pgen.1007926. eCollection 2019 Jan. PLoS Genet. 2019. PMID: 30677014 Free PMC article.
-
Arrested development: coordinating regeneration with development and growth in Drosophila melanogaster.Curr Opin Genet Dev. 2016 Oct;40:87-94. doi: 10.1016/j.gde.2016.06.008. Epub 2016 Jul 6. Curr Opin Genet Dev. 2016. PMID: 27394031 Free PMC article. Review.
-
Cell reprogramming during regeneration in Drosophila: transgression of compartment boundaries.Curr Opin Genet Dev. 2016 Oct;40:11-16. doi: 10.1016/j.gde.2016.05.008. Epub 2016 Jun 4. Curr Opin Genet Dev. 2016. PMID: 27266970 Review.
Cited by
-
An Open Question: Is Non-Ionizing Radiation a Tool for Controlling Apoptosis-Induced Proliferation?Int J Mol Sci. 2021 Oct 16;22(20):11159. doi: 10.3390/ijms222011159. Int J Mol Sci. 2021. PMID: 34681819 Free PMC article. Review.
-
Two-Faced: Roles of JNK Signalling During Tumourigenesis in the Drosophila Model.Front Cell Dev Biol. 2020 Feb 5;8:42. doi: 10.3389/fcell.2020.00042. eCollection 2020. Front Cell Dev Biol. 2020. PMID: 32117973 Free PMC article. Review.
-
Shared enhancer gene regulatory networks between wound and oncogenic programs.Elife. 2023 May 3;12:e81173. doi: 10.7554/eLife.81173. Elife. 2023. PMID: 37133250 Free PMC article.
-
Physiological and molecular mechanisms of insect appendage regeneration.Cell Regen. 2023 Mar 2;12(1):9. doi: 10.1186/s13619-022-00156-1. Cell Regen. 2023. PMID: 36859631 Free PMC article. Review.
-
Epithelial cell competition is promoted by signaling from immune cells.Nat Commun. 2025 Apr 19;16(1):3710. doi: 10.1038/s41467-025-59130-5. Nat Commun. 2025. PMID: 40251197 Free PMC article.
References
-
- Tanaka EM, Reddien PW. The cellular basis for animal regeneration. Dev Cell. 2011. July 19;21(1):172–85. doi: 10.1016/j.devcel.2011.06.016 - DOI - PMC - PubMed
-
- Hobmayer B, Rentzsch F, Kuhn K, Happel CM, Laue von CC, Snyder P, et al. WNT signalling molecules act in axis formation in the diploblastic metazoan Hydra. Nature. 2000. September 14;407(6801):186–9. doi: 10.1038/35025063 - DOI - PubMed
-
- Kawakami Y, Rodriguez Esteban C, Raya M, Kawakami H, Martí M, Dubova I, et al. Wnt/beta-catenin signaling regulates vertebrate limb regeneration. Genes Dev. 2006. December 1;20(23):3232–7. doi: 10.1101/gad.1475106 - DOI - PMC - PubMed
-
- Petersen CP, Reddien PW. Smed-betacatenin-1 is required for anteroposterior blastema polarity in planarian regeneration. Science. 2008. January 18;319(5861):327–30. doi: 10.1126/science.1149943 - DOI - PubMed
-
- Petersen CP, Reddien PW. A wound-induced Wnt expression program controls planarian regeneration polarity. Proc Natl Acad Sci USA. 2009. October 6;106(40):17061–6. doi: 10.1073/pnas.0906823106 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

