AID Recognizes Structured DNA for Class Switch Recombination
- PMID: 28757211
- PMCID: PMC5771415
- DOI: 10.1016/j.molcel.2017.06.034
AID Recognizes Structured DNA for Class Switch Recombination
Abstract
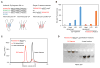
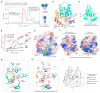
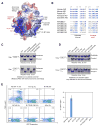
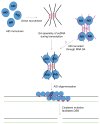
Activation-induced cytidine deaminase (AID) initiates both class switch recombination (CSR) and somatic hypermutation (SHM) in antibody diversification. Mechanisms of AID targeting and catalysis remain elusive despite its critical immunological roles and off-target effects in tumorigenesis. Here, we produced active human AID and revealed its preferred recognition and deamination of structured substrates. G-quadruplex (G4)-containing substrates mimicking the mammalian immunoglobulin switch regions are particularly good AID substrates in vitro. By solving crystal structures of maltose binding protein (MBP)-fused AID alone and in complex with deoxycytidine monophosphate, we surprisingly identify a bifurcated substrate-binding surface that explains structured substrate recognition by capturing two adjacent single-stranded overhangs simultaneously. Moreover, G4 substrates induce cooperative AID oligomerization. Structure-based mutations that disrupt bifurcated substrate recognition or oligomerization both compromise CSR in splenic B cells. Collectively, our data implicate intrinsic preference of AID for structured substrates and uncover the importance of G4 recognition and oligomerization of AID in CSR.
Keywords: AID; APOBEC; CSR; G-quadruplex; activation-induced cytidine deaminase; class switch recombination; crystal structure.
Copyright © 2017 Elsevier Inc. All rights reserved.
Figures



Similar articles
-
Non-redundancy of cytidine deaminases in class switch recombination.Eur J Immunol. 2004 Mar;34(3):844-849. doi: 10.1002/eji.200324418. Eur J Immunol. 2004. PMID: 14991614
-
Activation-induced deaminase (AID)-directed hypermutation in the immunoglobulin Smu region: implication of AID involvement in a common step of class switch recombination and somatic hypermutation.J Exp Med. 2002 Feb 18;195(4):529-34. doi: 10.1084/jem.20012144. J Exp Med. 2002. PMID: 11854365 Free PMC article.
-
DNA/RNA hybrid substrates modulate the catalytic activity of purified AID.Mol Immunol. 2018 Jan;93:94-106. doi: 10.1016/j.molimm.2017.11.012. Epub 2017 Nov 20. Mol Immunol. 2018. PMID: 29161581
-
Current insights into the mechanism of mammalian immunoglobulin class switch recombination.Crit Rev Biochem Mol Biol. 2019 Aug;54(4):333-351. doi: 10.1080/10409238.2019.1659227. Epub 2019 Sep 11. Crit Rev Biochem Mol Biol. 2019. PMID: 31509023 Free PMC article. Review.
-
Activation-induced cytidine deaminase: a dual role in class-switch recombination and somatic hypermutation.Eur J Immunol. 2003 Aug;33(8):2069-73. doi: 10.1002/eji.200324133. Eur J Immunol. 2003. PMID: 12884279 Review.
Cited by
-
AFF3, a susceptibility factor for autoimmune diseases, is a molecular facilitator of immunoglobulin class switch recombination.Sci Adv. 2022 Aug 26;8(34):eabq0008. doi: 10.1126/sciadv.abq0008. Epub 2022 Aug 24. Sci Adv. 2022. PMID: 36001653 Free PMC article.
-
Mutations at BCL11B Exon 4 Associated with T Cell Acute Lymphoblastic Leukemia Are Facilitated by AID and Formation of Non-B DNA Conformations.Mol Cell Biol. 2024;44(12):590-606. doi: 10.1080/10985549.2024.2419661. Epub 2024 Nov 7. Mol Cell Biol. 2024. PMID: 39511874
-
Nucleotide level mapping of uracils in murine heavy chain switch regions shows correlation between uracilation and positions of switch junctions created during class-switch recombination.NAR Mol Med. 2025 Feb 28;2(1):ugaf003. doi: 10.1093/narmme/ugaf003. eCollection 2025 Jan. NAR Mol Med. 2025. PMID: 40083807 Free PMC article.
-
The structure of APOBEC1 and insights into its RNA and DNA substrate selectivity.NAR Cancer. 2020 Dec;2(4):zcaa027. doi: 10.1093/narcan/zcaa027. Epub 2020 Oct 9. NAR Cancer. 2020. PMID: 33094286 Free PMC article.
-
Functions and consequences of AID/APOBEC-mediated DNA and RNA deamination.Nat Rev Genet. 2022 Aug;23(8):505-518. doi: 10.1038/s41576-022-00459-8. Epub 2022 Mar 7. Nat Rev Genet. 2022. PMID: 35256818 Free PMC article. Review.
References
-
- Basu U, Chaudhuri J, Alpert C, Dutt S, Ranganath S, Li G, Schrum JP, Manis JP, Alt FW. The AID antibody diversification enzyme is regulated by protein kinase A phosphorylation. Nature. 2005;438:508–511. - PubMed
-
- Chambers VS, Marsico G, Boutell JM, Di Antonio M, Smith GP, Balasubramanian S. High-throughput sequencing of DNA G-quadruplex structures in the human genome. Nat Biotechnol. 2015;33:877–881. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

