Ribosome Rescue Inhibitors Kill Actively Growing and Nonreplicating Persister Mycobacterium tuberculosis Cells
- PMID: 28762275
- PMCID: PMC5594445
- DOI: 10.1021/acsinfecdis.7b00028
Ribosome Rescue Inhibitors Kill Actively Growing and Nonreplicating Persister Mycobacterium tuberculosis Cells
Abstract
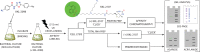
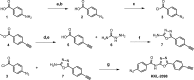
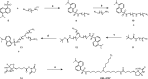
The emergence of Mycobacterium tuberculosis (MTB) strains that are resistant to most or all available antibiotics has created a severe problem for treating tuberculosis and has spurred a quest for new antibiotic targets. Here, we demonstrate that trans-translation is essential for growth of MTB and is a viable target for development of antituberculosis drugs. We also show that an inhibitor of trans-translation, KKL-35, is bactericidal against MTB under both aerobic and anoxic conditions. Biochemical experiments show that this compound targets helix 89 of the 23S rRNA. In silico molecular docking predicts a binding pocket for KKL-35 adjacent to the peptidyl-transfer center in a region not targeted by conventional antibiotics. Computational solvent mapping suggests that this pocket is a druggable hot spot for small molecule binding. Collectively, our findings reveal a new target for antituberculosis drug development and provide critical insight on the mechanism of antibacterial action for KKL-35 and related 1,3,4-oxadiazole benzamides.
Keywords: 1,3,4-oxadiazoles; Mycobacterium tuberculosis; antibiotics; ribosome rescue.
Conflict of interest statement
The authors declare no competing financial interest.
Figures





Similar articles
-
KKL-35 Exhibits Potent Antibiotic Activity against Legionella Species Independently of trans-Translation Inhibition.Antimicrob Agents Chemother. 2018 Jan 25;62(2):e01459-17. doi: 10.1128/AAC.01459-17. Print 2018 Feb. Antimicrob Agents Chemother. 2018. PMID: 29158279 Free PMC article.
-
Discovery of novel antituberculosis agents among 3-phenyl-5-(1-phenyl-1H-[1,2,3]triazol-4-yl)-[1,2,4]oxadiazole derivatives targeting aminoacyl-tRNA synthetases.Sci Rep. 2021 Mar 30;11(1):7162. doi: 10.1038/s41598-021-86562-y. Sci Rep. 2021. PMID: 33785838 Free PMC article.
-
In vitro anti-TB properties, in silico target validation, molecular docking and dynamics studies of substituted 1,2,4-oxadiazole analogues against Mycobacterium tuberculosis.J Enzyme Inhib Med Chem. 2021 Dec;36(1):869-884. doi: 10.1080/14756366.2021.1900162. J Enzyme Inhib Med Chem. 2021. PMID: 34060396 Free PMC article.
-
Systematic Review on 1,2,3-Oxadiazoles, 1,2,4-Oxadiazoles, and 1,2,5-Oxadiazoles in the Antimycobacterial Drug Discovery.ChemMedChem. 2025 May 5;20(9):e202400971. doi: 10.1002/cmdc.202400971. Epub 2025 Feb 4. ChemMedChem. 2025. PMID: 39846226
-
Oxadiazole scaffolds in anti-tuberculosis drug discovery.Bioorg Med Chem Lett. 2019 Aug 15;29(16):1999-2007. doi: 10.1016/j.bmcl.2019.06.054. Epub 2019 Jun 28. Bioorg Med Chem Lett. 2019. PMID: 31296357 Review.
Cited by
-
Tag-Dependent Substrate Selection of ClpX Underlies Secondary Differentiation of Chlamydia trachomatis.mBio. 2022 Oct 26;13(5):e0185822. doi: 10.1128/mbio.01858-22. Epub 2022 Sep 26. mBio. 2022. PMID: 36154190 Free PMC article.
-
The Structural and Molecular Mechanisms of Mycobacterium tuberculosis Translational Elongation Factor Proteins.Molecules. 2024 Apr 29;29(9):2058. doi: 10.3390/molecules29092058. Molecules. 2024. PMID: 38731549 Free PMC article. Review.
-
Improved Antibacterial Activity of 1,3,4-Oxadiazole-Based Compounds That Restrict Staphylococcus aureus Growth Independent of LtaS Function.ACS Infect Dis. 2023 Nov 10;9(11):2141-2159. doi: 10.1021/acsinfecdis.3c00250. Epub 2023 Oct 13. ACS Infect Dis. 2023. PMID: 37828912 Free PMC article.
-
Trans-Translation Is an Appealing Target for the Development of New Antimicrobial Compounds.Microorganisms. 2021 Dec 21;10(1):3. doi: 10.3390/microorganisms10010003. Microorganisms. 2021. PMID: 35056452 Free PMC article. Review.
-
A New Mechanism for Ribosome Rescue Can Recruit RF1 or RF2 to Nonstop Ribosomes.mBio. 2018 Dec 18;9(6):e02436-18. doi: 10.1128/mBio.02436-18. mBio. 2018. PMID: 30563899 Free PMC article.
References
-
- World Health Organization. (2016) Global tuberculosis report, World Health Organization, Geneva, Switzerland, http://www.who.int/tb/publications/global_report/en/.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

