Protospacer Adjacent Motif-Induced Allostery Activates CRISPR-Cas9
- PMID: 28764328
- PMCID: PMC5905990
- DOI: 10.1021/jacs.7b05313
Protospacer Adjacent Motif-Induced Allostery Activates CRISPR-Cas9
Abstract
CRISPR-Cas9 is a genome editing technology with major impact in life sciences. In this system, the endonuclease Cas9 generates double strand breaks in DNA upon RNA-guided recognition of a complementary DNA sequence, which strictly requires the presence of a protospacer adjacent motif (PAM) next to the target site. Although PAM recognition is essential for cleavage, it is unknown whether and how PAM binding activates Cas9 for DNA cleavage at spatially distant sites. Here, we find evidence of a PAM-induced allosteric mechanism revealed by microsecond molecular dynamics simulations. PAM acts as an allosteric effector and triggers the interdependent conformational dynamics of the Cas9 catalytic domains (HNH and RuvC), responsible for concerted cleavage of the two DNA strands. Targeting such an allosteric mechanism should enable control of CRISPR-Cas9 functionality.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

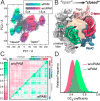

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

