Predicting Organ Toxicity Using in Vitro Bioactivity Data and Chemical Structure
- PMID: 28768096
- PMCID: PMC6172960
- DOI: 10.1021/acs.chemrestox.7b00084
Predicting Organ Toxicity Using in Vitro Bioactivity Data and Chemical Structure
Abstract
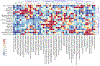
Animal testing alone cannot practically evaluate the health hazard posed by tens of thousands of environmental chemicals. Computational approaches making use of high-throughput experimental data may provide more efficient means to predict chemical toxicity. Here, we use a supervised machine learning strategy to systematically investigate the relative importance of study type, machine learning algorithm, and type of descriptor on predicting in vivo repeat-dose toxicity at the organ-level. A total of 985 compounds were represented using chemical structural descriptors, ToxPrint chemotype descriptors, and bioactivity descriptors from ToxCast in vitro high-throughput screening assays. Using ToxRefDB, a total of 35 target organ outcomes were identified that contained at least 100 chemicals (50 positive and 50 negative). Supervised machine learning was performed using Naïve Bayes, k-nearest neighbor, random forest, classification and regression trees, and support vector classification approaches. Model performance was assessed based on F1 scores using 5-fold cross-validation with balanced bootstrap replicates. Fixed effects modeling showed the variance in F1 scores was explained mostly by target organ outcome, followed by descriptor type, machine learning algorithm, and interactions between these three factors. A combination of bioactivity and chemical structure or chemotype descriptors were the most predictive. Model performance improved with more chemicals (up to a maximum of 24%), and these gains were correlated (ρ = 0.92) with the number of chemicals. Overall, the results demonstrate that a combination of bioactivity and chemical descriptors can accurately predict a range of target organ toxicity outcomes in repeat-dose studies, but specific experimental and methodologic improvements may increase predictivity.
Figures




References
-
- Wagner K, Fach B, and Kolar R (2012) Inconsistencies in data requirements of EU legislation involving tests on animals. ALTEX 29, 302–332. - PubMed
-
- Everts S (2009) Cost Of REACH Underestimated. Chemical and Engineering News 87, 7.
-
- EPA. (2016) TSCA Chemical Substance Inventory, U.S. Environmental Protection Agency.
-
- ECHA. (2016) European Chemicals Agency (ECHA): Pre-registered Substances. Pre-registered substances - ECHA.
-
- Rovida C, and Hartung T (2009) Re-evaluation of animal numbers and costs for in vivo tests to accomplish REACH legislation requirements for chemicals - a report by the transatlantic think tank for toxicology (t(4)). ALTEX 26, 187–208. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

