Structural and mutational analyses of the Leptospira interrogans virulence-related heme oxygenase provide insights into its catalytic mechanism
- PMID: 28771589
- PMCID: PMC5542595
- DOI: 10.1371/journal.pone.0182535
Structural and mutational analyses of the Leptospira interrogans virulence-related heme oxygenase provide insights into its catalytic mechanism
Abstract
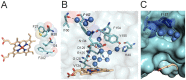
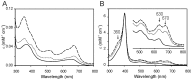
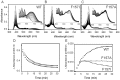
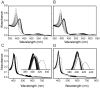
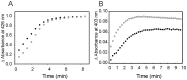
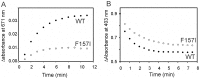
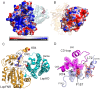
Heme oxygenase from Leptospira interrogans is an important virulence factor. During catalysis, redox equivalents are provided to this enzyme by the plastidic-type ferredoxin-NADP+ reductase also found in L. interrogans. This process may have evolved to aid this bacterial pathogen to obtain heme-iron from their host and enable successful colonization. Herein we report the crystal structure of the heme oxygenase-heme complex at 1.73 Å resolution. The structure reveals several distinctive features related to its function. A hydrogen bonded network of structural water molecules that extends from the catalytic site to the protein surface was cleared observed. A depression on the surface appears to be the H+ network entrance from the aqueous environment to the catalytic site for O2 activation, a key step in the heme oxygenase reaction. We have performed a mutational analysis of the F157, located at the above-mentioned depression. The mutant enzymes were unable to carry out the complete degradation of heme to biliverdin since the reaction was arrested at the verdoheme stage. We also observed that the stability of the oxyferrous complex, the efficiency of heme hydroxylation and the subsequent conversion to verdoheme was adversely affected. These findings underscore a long-range communication between the outer fringes of the hydrogen-bonded network of structural waters and the heme active site during catalysis. Finally, by analyzing the crystal structures of ferredoxin-NADP+ reductase and heme oxygenase, we propose a model for the productive association of these proteins.
Conflict of interest statement
Figures

References
-
- Benson DR, Rivera M. Heme uptake and metabolism in bacteria. Met Ions Life Sci. 2013;12: 279–332. doi: 10.1007/978-94-007-5561-1_9 - DOI - PubMed
-
- Rivera M, Zeng Y. Heme oxygenase, steering dioxygen activation toward heme hydroxylation. J Inorg Biochem. 2005;99: 337–54. doi: 10.1016/j.jinorgbio.2004.09.016 - DOI - PubMed
-
- Rivera M, Rodríguez JC. The dual role of heme as cofactor and substrate in the biosynthesis of carbon monoxide. Met Ions Life Sci. 2009;6: 241–93. doi: 10.1039/BK9781847559159-00241 - DOI - PubMed
-
- Matsui T, Unno M, Ikeda-saito M. Heme Oxygenase Reveals Its Strategy for Catalyzing Three Successive Oxygenation Reactions. Acc Chem Res. 2010;43: 240–247. doi: 10.1021/ar9001685 - DOI - PubMed
-
- Wilks A. Heme Oxygenase: Evolution, Structure, and Mechanism. Antioxid Redox Signal. 2002;4: 603–614. doi: 10.1089/15230860260220102 - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

