UBE2O remodels the proteome during terminal erythroid differentiation
- PMID: 28774900
- PMCID: PMC5812729
- DOI: 10.1126/science.aan0218
UBE2O remodels the proteome during terminal erythroid differentiation
Abstract
During terminal differentiation, the global protein complement is remodeled, as epitomized by erythrocytes, whose cytosol is ~98% globin. The erythroid proteome undergoes a rapid transition at the reticulocyte stage; however, the mechanisms driving programmed elimination of preexisting cytosolic proteins are unclear. We found that a mutation in the murine Ube2o gene, which encodes a ubiquitin-conjugating enzyme induced during erythropoiesis, results in anemia. Proteomic analysis suggested that UBE2O is a broad-spectrum ubiquitinating enzyme that remodels the erythroid proteome. In particular, ribosome elimination, a hallmark of reticulocyte differentiation, was defective in Ube2o-/- mutants. UBE2O recognized ribosomal proteins and other substrates directly, targeting them to proteasomes for degradation. Thus, in reticulocytes, the induction of ubiquitinating factors may drive the transition from a complex to a simple proteome.
Copyright © 2017 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Figures
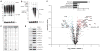
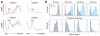



Comment in
-
New developments for protein quality control.Science. 2017 Aug 4;357(6350):450-451. doi: 10.1126/science.aao1896. Science. 2017. PMID: 28774913 No abstract available.
References
-
- Roux-Dalvai F, et al. Extensive analysis of the cytoplasmic proteome of human erythrocytes using the peptide ligand library technology and advanced mass spectrometry. Mol Cell Proteomics. 2008;7:2254–2269. - PubMed
-
- Hass AL. In: Red blood cell aging. Magnani M, De Flora A, editors. Plenum Press; New York and London: 1991. pp. 191–205.
-
- Ciehanover A, Hod Y, Hershko A. A heat-stable polypeptide component of an ATP-dependent proteolytic system from reticulocytes. Biochem Biophys Res Commun. 1978;81:1100–1105. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

