Divergent co-transcriptomes of different host cells infected with Toxoplasma gondii reveal cell type-specific host-parasite interactions
- PMID: 28775382
- PMCID: PMC5543063
- DOI: 10.1038/s41598-017-07838-w
Divergent co-transcriptomes of different host cells infected with Toxoplasma gondii reveal cell type-specific host-parasite interactions
Abstract
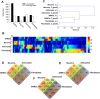
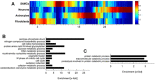
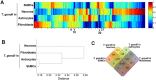
The apicomplexan parasite Toxoplasma gondii infects various cell types in avian and mammalian hosts including humans. Infection of immunocompetent hosts is mostly asymptomatic or benign, but leads to development of largely dormant bradyzoites that persist predominantly within neurons and muscle cells. Here we have analyzed the impact of the host cell type on the co-transcriptomes of host and parasite using high-throughput RNA sequencing. Murine cortical neurons and astrocytes, skeletal muscle cells (SkMCs) and fibroblasts differed by more than 16,200 differentially expressed genes (DEGs) before and after infection with T. gondii. However, only a few hundred of them were regulated by infection and these largely diverged in neurons, SkMCs, astrocytes and fibroblasts indicating host cell type-specific transcriptional responses after infection. The heterogeneous transcriptomes of host cells before and during infection coincided with ~5,400 DEGs in T. gondii residing in different cell types. Finally, we identified gene clusters in both T. gondii and its host, which correlated with the predominant parasite persistence in neurons or SkMCs as compared to astrocytes or fibroblasts. Thus, heterogeneous expression profiles of different host cell types and the parasites' ability to adapting to them may govern the parasite-host cell interaction during toxoplasmosis.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

