Genome-wide signatures of complex introgression and adaptive evolution in the big cats
- PMID: 28776029
- PMCID: PMC5517113
- DOI: 10.1126/sciadv.1700299
Genome-wide signatures of complex introgression and adaptive evolution in the big cats
Abstract
The great cats of the genus Panthera comprise a recent radiation whose evolutionary history is poorly understood. Their rapid diversification poses challenges to resolving their phylogeny while offering opportunities to investigate the historical dynamics of adaptive divergence. We report the sequence, de novo assembly, and annotation of the jaguar (Panthera onca) genome, a novel genome sequence for the leopard (Panthera pardus), and comparative analyses encompassing all living Panthera species. Demographic reconstructions indicated that all of these species have experienced variable episodes of population decline during the Pleistocene, ultimately leading to small effective sizes in present-day genomes. We observed pervasive genealogical discordance across Panthera genomes, caused by both incomplete lineage sorting and complex patterns of historical interspecific hybridization. We identified multiple signatures of species-specific positive selection, affecting genes involved in craniofacial and limb development, protein metabolism, hypoxia, reproduction, pigmentation, and sensory perception. There was remarkable concordance in pathways enriched in genomic segments implicated in interspecies introgression and in positive selection, suggesting that these processes were connected. We tested this hypothesis by developing exome capture probes targeting ~19,000 Panthera genes and applying them to 30 wild-caught jaguars. We found at least two genes (DOCK3 and COL4A5, both related to optic nerve development) bearing significant signatures of interspecies introgression and within-species positive selection. These findings indicate that post-speciation admixture has contributed genetic material that facilitated the adaptive evolution of big cat lineages.
Figures
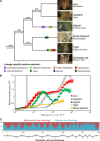



References
-
- J. A. Coyne, H. A. Orr, Speciation (Sinauer Associates, 2004).
-
- Fontaine M. C., Pease J. B., Steele A., Waterhouse R. M., Neafsey D. E., Sharakhov I. V., Jiang X., Hall A. B., Catteruccia F., Kakani E., Mitchell S. N., Wu Y.-C., Smith H. A., Love R. R., Lawniczak M. K., Slotman M. A., Emrich S. J., Hahn M. W., Besansky N. J., Extensive introgression in a malaria vector species complex revealed by phylogenomics. Science 347, 1258524 (2015). - PMC - PubMed
-
- Lamichhaney S., Berglund J., Almén M. S., Maqbool K., Grabherr M., Martinez-Barrio A., Promerová M., Rubin C.-J., Wang C., Zamani N., Grant B. R., Grant P. R., Webster M. T., Andersson L., Evolution of Darwin’s finches and their beaks revealed by genome sequencing. Nature 518, 371–375 (2015). - PubMed
-
- Johnson W. E., Eizirik E., Pecon-Slattery J., Murphy W. J., Antunes A., Teeling E., O’Brien S. J., The late miocene radiation of modern felidae: A genetic assessment. Science 311, 73–77 (2006). - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

