An efficient strategy using k- mers to analyse 16S rRNA sequences
- PMID: 28795166
- PMCID: PMC5537200
- DOI: 10.1016/j.heliyon.2017.e00370
An efficient strategy using k- mers to analyse 16S rRNA sequences
Abstract
The use of k-mers has been a successful strategy for improving metagenomics studies, including taxonomic classifications, or de novo assemblies, and can be used to obtain sequences of interest from the available databases. The aim of this manuscript was to propose a simple but efficient strategy to generate k-mers and to use them to obtain and analyse in silico 16S rRNA sequence fragments. A total of 513,309 bacterial sequences contained in the SILVA database were considered for the study, and homemade PHP scripts were used to search for specific nucleotide chains, recover fragments of bacterial sequences, make calculations and organize information. Consensus sequences matching conserved regions were constructed by aligning most of the primers used in the literature. Sequences of k nucleotides (9- to 15-mers) were extracted from the generated primer contigs. Frequency analysis revealed that k-mer size was inversely proportional to the occurrence of k-mers in the different conserved regions, suggesting a stringency relationship; high numbers of duplicate reactions were observed with short k-mers, and a lower proportion of sequences were obtained with large ones, with the best results obtained using 12-mers. Using 12-mers with the proposed method to obtain and study sequences was found to be a reliable approach for the analysis of 16S rRNA sequences and this strategy may probably be extended to other biomarkers. Furthermore, additional applications such as evaluating the degree of conservation and designing primers and other calculations are proposed as examples.
Keywords: Bioinformatics; Biological sciences; Microbiology.
Figures
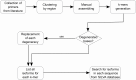




References
-
- Pandey S., Singh S., Yadav A.N., Nain L., Saxena A.K. Phylogenetic diversity and characterization of novel and efficient cellulase producing bacterial isolates from various extreme environments. Biosci. Biotechnol. Biochem. 2013;77:1474–1480. - PubMed
-
- Lakaniemi A.-M., Hulatt C.J., Wakeman K.D., Thomas D.N., Puhakka J.A. Eukaryotic and prokaryotic microbial communities during microalgal biomass production. Bioresour. Technol. 2012;124:387–393. - PubMed
-
- Stackebrandt E., Goebel B. Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int. J. Syst. Evol. Microbiol. 1994;44:846–849.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

