Identification of key lncRNAs in colorectal cancer progression based on associated protein-protein interaction analysis
- PMID: 28797257
- PMCID: PMC5553992
- DOI: 10.1186/s12957-017-1211-7
Identification of key lncRNAs in colorectal cancer progression based on associated protein-protein interaction analysis
Abstract
Background: Colorectal cancer (CRC) was one of the most commonly diagnosed malignancies. The molecular mechanisms involved in the progression of CRC remain unclear. Accumulating evidences showed that long noncoding RNAs (lncRNAs) played key roles in tumorigenesis, cancer progression, and metastasis. Therefore, we aimed to explore the roles of lncRNAs in the progression of CRC.
Methods: In this study, we aimed to identify differentially expressed lncRNAs and messenger RNAs (mRNAs) in CRC by analyzing a cohort of previously published datasets: GSE64857. GO and KEGG pathway analyses were applied to give us insight in the functions of those lncRNAs and mRNAs in CRC.
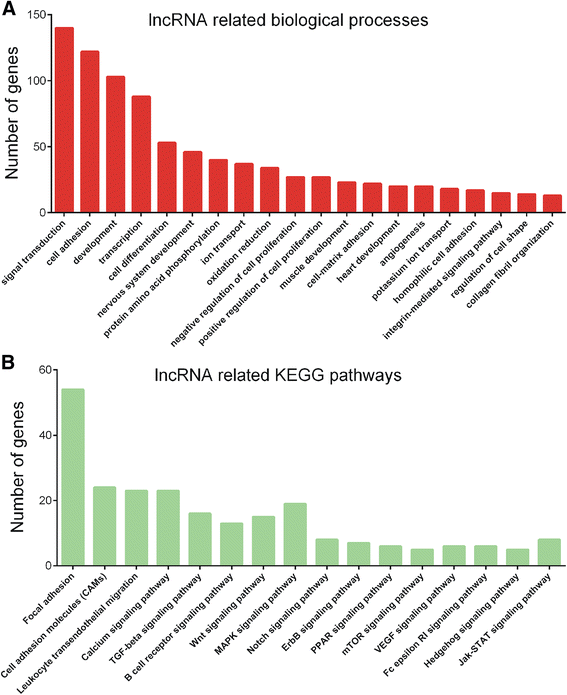
Results: Totally, 46 lncRNAs were identified as differentially expressed between stage II and stage III CRC for the first time screening by microarray. GO and KEGG pathway analyses showed that differentially expressed lncRNAs were involved in regulating signal transduction, cell adhesion, cell differentiation, focal adhesion, and cell adhesion molecules.
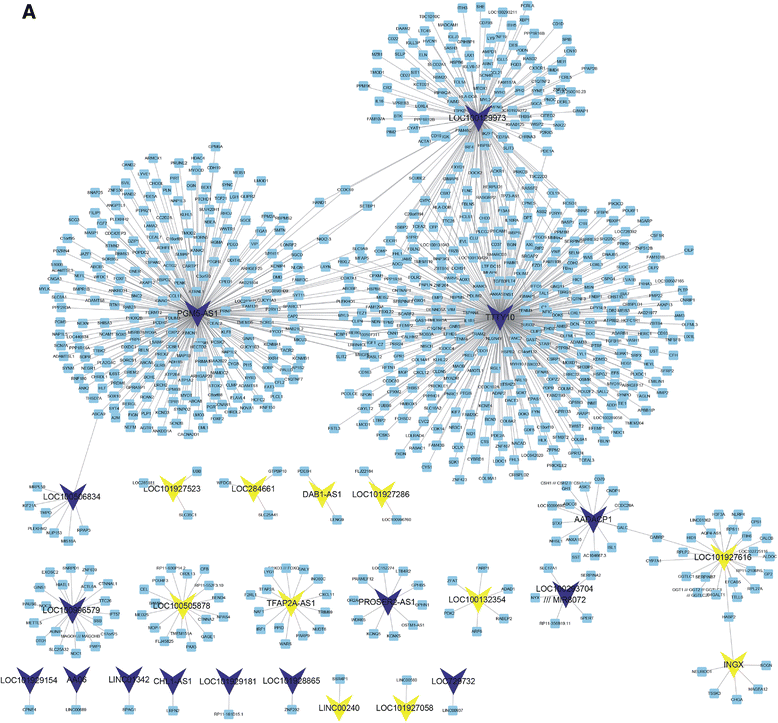
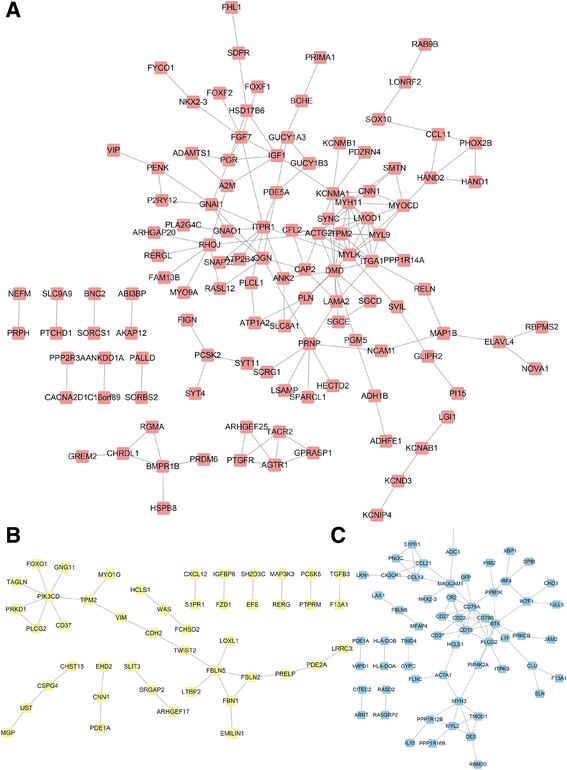
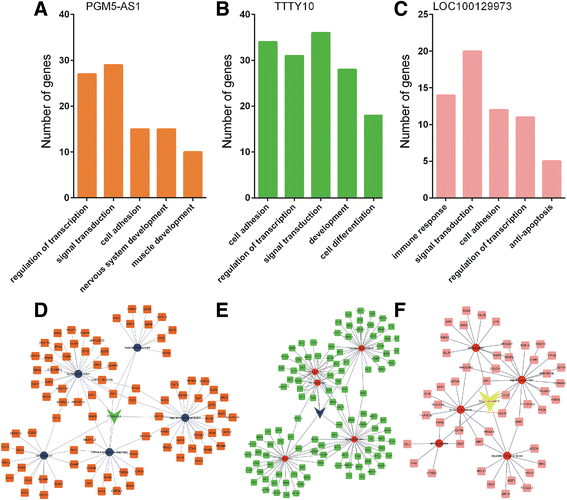
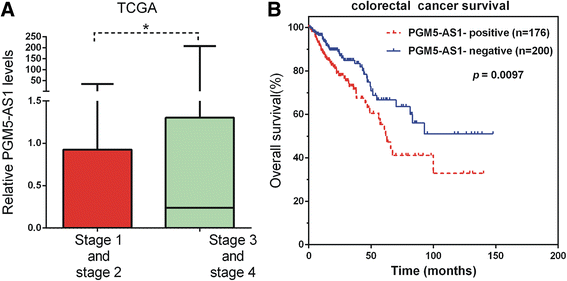
Conclusions: We found three lncRNAs (LOC100129973, PGM5-AS1, and TTTY10) widely co-expressed with differentially expressed mRNAs. We also constructed lncRNA-associated PPI in CRC and found that these lncRNAs may be associated with CRC progression. Moreover, we found that high PGM5-AS1 expression levels were associated with worse overall survival in CRC cancer. We believe that this study would provide novel potential therapeutic and prognostic targets for CRC.
Keywords: Colorectal cancer; Expression profiling; Long non-coding RNA; Protein–protein interaction analysis.
Conflict of interest statement
Competing interest
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
Similar articles
-
Comprehensive analysis of the long noncoding RNA expression profile and construction of the lncRNA-mRNA co-expression network in colorectal cancer.Cancer Biol Ther. 2020;21(2):157-169. doi: 10.1080/15384047.2019.1673098. Epub 2019 Oct 16. Cancer Biol Ther. 2020. PMID: 31619123 Free PMC article.
-
Overexpression of lncRNA AFAP1-AS1 correlates with poor prognosis and promotes tumorigenesis in colorectal cancer.Biomed Pharmacother. 2016 Jul;81:152-159. doi: 10.1016/j.biopha.2016.04.009. Epub 2016 Apr 16. Biomed Pharmacother. 2016. PMID: 27261589
-
Long noncoding RNA HNF1A-AS1 indicates a poor prognosis of colorectal cancer and promotes carcinogenesis via activation of the Wnt/β-catenin signaling pathway.Biomed Pharmacother. 2017 Dec;96:877-883. doi: 10.1016/j.biopha.2017.10.033. Epub 2017 Nov 6. Biomed Pharmacother. 2017. PMID: 29145164
-
Identification of MFI2-AS1, a Novel Pivotal lncRNA for Prognosis of Stage III/IV Colorectal Cancer.Dig Dis Sci. 2020 Dec;65(12):3538-3550. doi: 10.1007/s10620-020-06064-1. Epub 2020 Jan 20. Dig Dis Sci. 2020. PMID: 31960204 Review.
-
The Biological Effect and Clinical Application of Long Noncoding RNAs in Colorectal Cancer.Cell Physiol Biochem. 2018;46(2):431-441. doi: 10.1159/000488610. Epub 2018 Mar 26. Cell Physiol Biochem. 2018. PMID: 29614491 Review.
Cited by
-
Insulin-Like Growth Factor 2 (IGF2) Signaling in Colorectal Cancer-From Basic Research to Potential Clinical Applications.Int J Mol Sci. 2019 Oct 3;20(19):4915. doi: 10.3390/ijms20194915. Int J Mol Sci. 2019. PMID: 31623387 Free PMC article. Review.
-
Knockdown of lncRNA MIAT inhibits proliferation and cisplatin resistance in non-small cell lung cancer cells by increasing miR-184 expression.Oncol Lett. 2020 Jan;19(1):533-541. doi: 10.3892/ol.2019.11084. Epub 2019 Nov 13. Oncol Lett. 2020. PMID: 31897168 Free PMC article.
-
More than the SRY: The Non-Coding Landscape of the Y Chromosome and Its Importance in Human Disease.Noncoding RNA. 2024 Apr 10;10(2):21. doi: 10.3390/ncrna10020021. Noncoding RNA. 2024. PMID: 38668379 Free PMC article. Review.
-
Long non-coding RNA profiling of pediatric Medulloblastoma.BMC Med Genomics. 2020 Jun 26;13(1):87. doi: 10.1186/s12920-020-00744-7. BMC Med Genomics. 2020. PMID: 32591022 Free PMC article.
-
LncRNA ZEB2-AS1 contributes to the tumorigenesis of gastric cancer via activating the Wnt/β-catenin pathway.Mol Cell Biochem. 2019 Jun;456(1-2):73-83. doi: 10.1007/s11010-018-03491-7. Epub 2019 Jan 11. Mol Cell Biochem. 2019. PMID: 30635820
References
-
- Peng W, Wang Z, Fan H. LncRNA NEAT1 impacts cell proliferation and apoptosis of colorectal cancer via regulation of Akt signaling. Pathol Oncol Res. 2016;23(3):651-56. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

