Original Chemical Series of Pyrimidine Biosynthesis Inhibitors That Boost the Antiviral Interferon Response
- PMID: 28807907
- PMCID: PMC5610480
- DOI: 10.1128/AAC.00383-17
Original Chemical Series of Pyrimidine Biosynthesis Inhibitors That Boost the Antiviral Interferon Response
Abstract
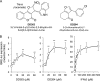
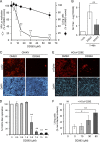
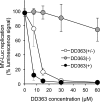
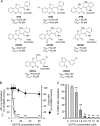
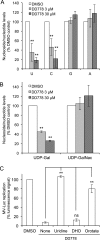
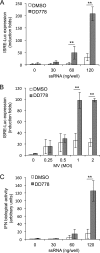
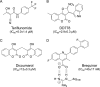
De novo pyrimidine biosynthesis is a key metabolic pathway involved in multiple biosynthetic processes. Here, we identified an original series of 3-(1H-indol-3-yl)-2,3-dihydro-4H-furo[3,2-c]chromen-4-one derivatives as a new class of pyrimidine biosynthesis inhibitors formed by two edge-fused polycyclic moieties. We show that identified compounds exhibit broad-spectrum antiviral activity and immunostimulatory properties, in line with recent reports linking de novo pyrimidine biosynthesis with innate defense mechanisms against viruses. Most importantly, we establish that pyrimidine deprivation can amplify the production of both type I and type III interferons by cells stimulated with retinoic acid-inducible gene 1 (RIG-I) ligands. Altogether, our results further expand the current panel of pyrimidine biosynthesis inhibitors and illustrate how the production of antiviral interferons is tightly coupled to this metabolic pathway. Functional and structural similarities between this new chemical series and dicoumarol, which was reported before to inhibit pyrimidine biosynthesis at the dihydroorotate dehydrogenase (DHODH) step, are discussed.
Keywords: antiviral agents; innate immunity; interferons; pyrimidine metabolism.
Copyright © 2017 Lucas-Hourani et al.
Figures


References
-
- Bonavia A, Franti M, Pusateri Keaney E, Kuhen K, Seepersaud M, Radetich B, Shao J, Honda A, Dewhurst J, Balabanis K, Monroe J, Wolff K, Osborne C, Lanieri L, Hoffmaster K, Amin J, Markovits J, Broome M, Skuba E, Cornella-Taracido I, Joberty G, Bouwmeester T, Hamann L, Tallarico JA, Tommasi R, Compton T, Bushell SM. 2011. Identification of broad-spectrum antiviral compounds and assessment of the druggability of their target for efficacy against respiratory syncytial virus (RSV). Proc Natl Acad Sci U S A 108:6739–6744. doi: 10.1073/pnas.1017142108. - DOI - PMC - PubMed
-
- Wang QY, Bushell S, Qing M, Xu HY, Bonavia A, Nunes S, Zhou J, Poh MK, Florez de Sessions P, Niyomrattanakit P, Dong H, Hoffmaster K, Goh A, Nilar S, Schul W, Jones S, Kramer L, Compton T, Shi PY. 2011. Inhibition of dengue virus through suppression of host pyrimidine biosynthesis. J Virol 85:6548–6556. doi: 10.1128/JVI.02510-10. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

