Directly monitor protein rearrangement on a nanosecond-to-millisecond time-scale
- PMID: 28821738
- PMCID: PMC5562898
- DOI: 10.1038/s41598-017-08385-0
Directly monitor protein rearrangement on a nanosecond-to-millisecond time-scale
Abstract
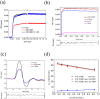
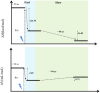
In order to directly observe the refolding kinetics from a partially misfolded state to a native state in the bottom of the protein-folding funnel, we used a "caging" strategy to trap the β-sheet structure of ubiquitin in a misfolded conformation. We used molecular dynamics simulation to generate the cage-induced, misfolded structure and compared the structure of the misfolded ubiquitin with native ubiquitin. Using laser flash irradiation, the cage can be cleaved from the misfolded structure within one nanosecond, and we monitored the refolding kinetics of ubiquitin from this misfolded state to the native state by photoacoustic calorimetry and photothermal beam deflection techniques on nanosecond to millisecond timescales. Our results showed two refolding events in this refolding process. The fast event is shorter than 20 ns and corresponds to the instant collapse of ubiquitin upon cage release initiated by laser irradiation. The slow event is ~60 μs, derived from a structural rearrangement in β-sheet refolding. The event lasts 10 times longer than the timescale of β-hairpin formation for short peptides as monitored by temperature jump, suggesting that rearrangement of a β-sheet structure from a misfolded state to its native state requires more time than ab initio folding of a β-sheet.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




Similar articles
-
Temperature-dependent downhill unfolding of ubiquitin. I. Nanosecond-to-millisecond resolved nonlinear infrared spectroscopy.Proteins. 2008 Jul;72(1):474-87. doi: 10.1002/prot.22043. Proteins. 2008. PMID: 18384151
-
Conformational changes during the nanosecond-to-millisecond unfolding of ubiquitin.Proc Natl Acad Sci U S A. 2005 Jan 18;102(3):612-7. doi: 10.1073/pnas.0408646102. Epub 2005 Jan 3. Proc Natl Acad Sci U S A. 2005. PMID: 15630083 Free PMC article.
-
The interplay of turn formation and hydrophobic interactions on the early kinetic events in protein folding.Chem Commun (Camb). 2012 Jan 14;48(4):487-97. doi: 10.1039/c1cc13278d. Epub 2011 Nov 4. Chem Commun (Camb). 2012. PMID: 22053320
-
Measuring the refolding of beta-sheets with different turn sequences on a nanosecond time scale.Proc Natl Acad Sci U S A. 2004 May 11;101(19):7305-10. doi: 10.1073/pnas.0304922101. Epub 2004 May 3. Proc Natl Acad Sci U S A. 2004. PMID: 15123838 Free PMC article.
-
Simulating the peptide folding kinetic related spectra based on the Markov State Model.Adv Exp Med Biol. 2014;805:199-220. doi: 10.1007/978-3-319-02970-2_9. Adv Exp Med Biol. 2014. PMID: 24446363 Review.
Cited by
-
Interface integrity in septin protofilaments is maintained by an arginine residue conserved from yeast to man.Mol Biol Cell. 2025 May 1;36(5):ar59. doi: 10.1091/mbc.E25-01-0041. Epub 2025 Mar 26. Mol Biol Cell. 2025. PMID: 40137961
-
Protons Are Fast and Smart; Proteins Are Slow and Dumb: On the Relationship of Electrospray Ionization Charge States and Conformations.J Am Soc Mass Spectrom. 2021 Jul 7;32(7):1553-1561. doi: 10.1021/jasms.1c00100. Epub 2021 Jun 21. J Am Soc Mass Spectrom. 2021. PMID: 34151568 Free PMC article.
-
Entropy-Enthalpy Compensations Fold Proteins in Precise Ways.Int J Mol Sci. 2021 Sep 6;22(17):9653. doi: 10.3390/ijms22179653. Int J Mol Sci. 2021. PMID: 34502559 Free PMC article.
-
Visible-to-NIR-Light Activated Release: From Small Molecules to Nanomaterials.Chem Rev. 2020 Dec 23;120(24):13135-13272. doi: 10.1021/acs.chemrev.0c00663. Epub 2020 Oct 30. Chem Rev. 2020. PMID: 33125209 Free PMC article.
-
Characterization of the structural forces governing the reversibility of the thermal unfolding of the human acidic fibroblast growth factor.Sci Rep. 2021 Aug 2;11(1):15579. doi: 10.1038/s41598-021-95050-2. Sci Rep. 2021. PMID: 34341408 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

