Live-cell p53 single-molecule binding is modulated by C-terminal acetylation and correlates with transcriptional activity
- PMID: 28827596
- PMCID: PMC5567047
- DOI: 10.1038/s41467-017-00398-7
Live-cell p53 single-molecule binding is modulated by C-terminal acetylation and correlates with transcriptional activity
Abstract
Live-cell microscopy has highlighted that transcription factors bind transiently to chromatin but it is not clear if the duration of these binding interactions can be modulated in response to an activation stimulus, and if such modulation can be controlled by post-translational modifications of the transcription factor. We address this question for the tumor suppressor p53 by combining live-cell single-molecule microscopy and single cell in situ measurements of transcription and we show that p53-binding kinetics are modulated following genotoxic stress. The modulation of p53 residence times on chromatin requires C-terminal acetylation-a classical mark for transcriptionally active p53-and correlates with the induction of transcription of target genes such as CDKN1a. We propose a model in which the modification state of the transcription factor determines the coupling between transcription factor abundance and transcriptional activity by tuning the transcription factor residence time on target sites.Both transcription binding kinetics and post-translational modifications of transcription factors are thought to play a role in the modulation of transcription. Here the authors use single-molecule tracking to directly demonstrate that p53 acetylation modulates promoter residence time and transcriptional activity.
Conflict of interest statement
The authors declare no competing financial interests.
Figures
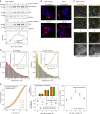




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

