NMR resonance assignments of the EVH1 domain of neurofibromin's recruitment factor Spred1
- PMID: 28831766
- PMCID: PMC5594049
- DOI: 10.1007/s12104-017-9768-1
NMR resonance assignments of the EVH1 domain of neurofibromin's recruitment factor Spred1
Abstract
Neurofibromin and Sprouty-related EVH1 domain-containing protein 1 (Spred1) both act as negative regulators of the mitogen-activated protein kinase pathway and are associated with the rare diseases Neurofibromatosis type 1 and Legius syndrome, respectively. Spred1 recruits the major GTPase activating protein (GAP) neurofibromin from the cytosol to the membrane in order to inactivate the small G protein Ras. These functions are dependent on the N-terminal EVH1 domain and the C-terminal Sprouty domain of Spred1 whereas the former specifically recognizes the GAP related domain of neurofibromin and the latter is responsible for membrane targeting. Within the GAP domain, Spred1 binding depends on the GAPex portion which is dispensable for Ras inactivation. In a first step towards the characterization of the Neurofibromin Spred1 interface in solution we assigned backbone and side chain 1H, 13C, and 15N chemical shifts of the Spred1 derived EVH1 domain. Our chemical shift data analysis indicate seven consecutive β-strands followed by a C-terminal α-helix which is in agreement with the previously reported crystal structure of Spred1(EVH1). Our data provide a framework for further analysis of the function of patient-derived mutations associated with rare diseases.
Keywords: Legius syndrome; NMR resonance assignment; Protein purification; TALOS+ prediction.
Figures
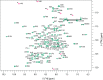

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

