Attenuated Human Parainfluenza Virus Type 1 Expressing the Respiratory Syncytial Virus (RSV) Fusion (F) Glycoprotein from an Added Gene: Effects of Prefusion Stabilization and Packaging of RSV F
- PMID: 28835504
- PMCID: PMC5660469
- DOI: 10.1128/JVI.01101-17
Attenuated Human Parainfluenza Virus Type 1 Expressing the Respiratory Syncytial Virus (RSV) Fusion (F) Glycoprotein from an Added Gene: Effects of Prefusion Stabilization and Packaging of RSV F
Abstract
Human respiratory syncytial virus (RSV) is the most prevalent worldwide cause of severe respiratory tract infection in infants and young children. Human parainfluenza virus type 1 (HPIV1) also causes severe pediatric respiratory illness, especially croup. Both viruses lack vaccines. Here, we describe the preclinical development of a bivalent RSV/HPIV1 vaccine based on a recombinant HPIV1 vector, attenuated by a stabilized mutation, that expresses RSV F protein modified for increased stability in the prefusion (pre-F) conformation by previously described disulfide bond (DS) and hydrophobic cavity-filling (Cav1) mutations. RSV F was expressed from the first or second gene position as the full-length protein or as a chimeric protein with its transmembrane and cytoplasmic tail (TMCT) domains substituted with those of HPIV1 F in an effort to direct packaging in the vector particles. All constructs were recovered by reverse genetics. The TMCT versions of RSV F were packaged in the rHPIV1 particles much more efficiently than their full-length counterparts. In hamsters, the presence of the RSV F gene, and in particular the TMCT versions, was attenuating and resulted in reduced immunogenicity. However, the vector expressing full-length RSV F from the pre-N position was immunogenic for RSV and HPIV1. It conferred complement-independent high-quality RSV-neutralizing antibodies at titers similar to those of wild-type RSV and provided protection against RSV challenge. The vectors exhibited stable RSV F expression in vitro and in vivo In conclusion, an attenuated rHPIV1 vector expressing a pre-F-stabilized form of RSV F demonstrated promising immunogenicity and should be further developed as an intranasal pediatric vaccine.IMPORTANCE RSV and HPIV1 are major viral causes of acute pediatric respiratory illness for which no vaccines or suitable antiviral drugs are available. The RSV F glycoprotein is the major RSV neutralization antigen. We used a rHPIV1 vector, bearing a stabilized attenuating mutation, to express the RSV F glycoprotein bearing amino acid substitutions that increase its stability in the pre-F form, the most immunogenic form that elicits highly functional virus-neutralizing antibodies. RSV F was expressed from the pre-N or N-P gene position of the rHPIV1 vector as a full-length protein or as a chimeric form with its TMCT domain derived from HPIV1 F. TMCT modification greatly increased packaging of RSV F into the vector particles but also increased vector attenuation in vivo, resulting in reduced immunogenicity. In contrast, full-length RSV F expressed from the pre-N position was immunogenic, eliciting complement-independent RSV-neutralizing antibodies and providing protection against RSV challenge.
Keywords: human parainfluenza virus type 1; intranasal vaccine; live attenuated vaccine; mucosal vaccines; pediatric vaccine; prefusion F; respiratory syncytial virus; vaccine.
Copyright © 2017 American Society for Microbiology.
Figures


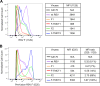
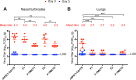


References
-
- Afonso CL, Amarasinghe GK, Banyai K, Bao Y, Basler CF, Bavari S, Bejerman N, Blasdell KR, Briand FX, Briese T, Bukreyev A, Calisher CH, Chandran K, Cheng J, Clawson AN, Collins PL, Dietzgen RG, Dolnik O, Domier LL, Durrwald R, Dye JM, Easton AJ, Ebihara H, Farkas SL, Freitas-Astua J, et al. . 2016. Taxonomy of the order Mononegavirales: update 2016. Arch Virol 161:2351–2360. doi:10.1007/s00705-016-2880-1. - DOI - PMC - PubMed
-
- Nair H, Nokes DJ, Gessner BD, Dherani M, Madhi SA, Singleton RJ, O'Brien KL, Roca A, Wright PF, Bruce N, Chandran A, Theodoratou E, Sutanto A, Sedyaningsih ER, Ngama M, Munywoki PK, Kartasasmita C, Simoes EA, Rudan I, Weber MW, Campbell H. 2010. Global burden of acute lower respiratory infections due to respiratory syncytial virus in young children: a systematic review and meta-analysis. Lancet 375:1545–1555. doi:10.1016/S0140-6736(10)60206-1. - DOI - PMC - PubMed
-
- Lozano R, Naghavi M, Foreman K, Lim S, Shibuya K, Aboyans V, Abraham J, Adair T, Aggarwal R, Ahn SY, Alvarado M, Anderson HR, Anderson LM, Andrews KG, Atkinson C, Baddour LM, Barker-Collo S, Bartels DH, et al. . 2012. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380:2095–2128. doi:10.1016/S0140-6736(12)61728-0. - DOI - PMC - PubMed
-
- Kapikian AZ, Mitchell RH, Chanock RM, Shvedoff RA, Stewart CE. 1969. An epidemiologic study of altered clinical reactivity to respiratory syncytial (RS) virus infection in children previously vaccinated with an inactivated RS virus vaccine. Am J Epidemiol 89:405–421. doi:10.1093/oxfordjournals.aje.a120954. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

