Rat Genome and Model Resources
- PMID: 28838068
- PMCID: PMC6057551
- DOI: 10.1093/ilar/ilw041
Rat Genome and Model Resources
Abstract
Rats remain a major model for studying disease mechanisms and discovery, validation, and testing of new compounds to improve human health. The rat's value continues to grow as indicated by the more than 1.4 million publications (second to human) at PubMed documenting important discoveries using this model. Advanced sequencing technologies, genome modification techniques, and the development of embryonic stem cell protocols ensure the rat remains an important mammalian model for disease studies. The 2004 release of the reference genome has been followed by the production of complete genomes for more than two dozen individual strains utilizing NextGen sequencing technologies; their analyses have identified over 80 million variants. This explosion in genomic data has been accompanied by the ability to selectively edit the rat genome, leading to hundreds of new strains through multiple technologies. A number of resources have been developed to provide investigators with access to precision rat models, comprehensive datasets, and sophisticated software tools necessary for their research. Those profiled here include the Rat Genome Database, PhenoGen, Gene Editing Rat Resource Center, Rat Resource and Research Center, and the National BioResource Project for the Rat in Japan.
Keywords: Rattus norvegicus; bioinformatics; database; disease; genomics; phenotype; rat; resource.
© The Author 2017. Published by Oxford University Press.
Figures



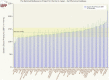
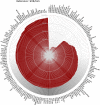

References
-
- Aken BL, Ayling S, Barrell D, Clarke L, Curwen V, Fairley S, Fernandez Banet J, Billis K, Garcia Giron C, Hourlier T, Howe K, Kahari A, Kokocinski F, Martin FJ, Murphy DN, Nag R, Ruffier M, Schuster M, Tang YA, Vogel JH, White S, Zadissa A, Flicek P, Searle SM. 2016. The Ensembl gene annotation system Database (Oxford) 2016. - PMC - PubMed
-
- Atanur SS, Diaz AG, Maratou K, Sarkis A, Rotival M, Game L, Tschannen MR, Kaisaki PJ, Otto GW, Ma MC, Keane TM, Hummel O, Saar K, Chen W, Guryev V, Gopalakrishnan K, Garrett MR, Joe B, Citterio L, Bianchi G, McBride M, Dominiczak A, Adams DJ, Serikawa T, Flicek P, Cuppen E, Hubner N, Petretto E, Gauguier D, Kwitek A, Jacob H, Aitman TJ. 2013. Genome sequencing reveals loci under artificial selection that underlie disease phenotypes in the laboratory rat. Cell 154(3):691–703. - PMC - PubMed
-
- Baud A, Hermsen R, Guryev V, Stridh P, Graham D, McBride MW, Foroud T, Calderari S, Diez M, Ockinger J, Beyeen AD, Gillett A, Abdelmagid N, Guerreiro-Cacais AO, Jagodic M, Tuncel J, Norin U, Beattie E, Huynh N, Miller WH, Koller DL, Alam I, Falak S, Osborne-Pellegrin M, Martinez-Membrives E, Canete T, Blazquez G, Vicens-Costa E, Mont-Cardona C, Diaz-Moran S, Tobena A, Hummel O, Zelenika D, Saar K, Patone G, Bauerfeind A, Bihoreau MT, Heinig M, Lee YA, Rintisch C, Schulz H, Wheeler DA, Worley KC, Muzny DM, Gibbs RA, Lathrop M, Lansu N, Toonen P, Ruzius FP, de Bruijn E, Hauser H, Adams DJ, Keane T, Atanur SS, Aitman TJ, Flicek P, Malinauskas T, Jones EY, Ekman D, Lopez-Aumatell R, Dominiczak AF, Johannesson M, Holmdahl R, Olsson T, Gauguier D, Hubner N, Fernandez-Teruel A, Cuppen E, Mott R, Flint J.. 2013. Combined sequence-based and genetic mapping analysis of complex traits in outbred rats. Nat Genet 45:767–775. - PMC - PubMed
-
- Bennett BJ, Farber CR, Orozco L, Kang HM, Ghazalpour A, Siemers N, Neubauer M, Neuhaus I, Yordanova R, Guan B, Truong A, Yang WP, He A, Kayne P, Gargalovic P, Kirchgessner T, Pan C, Castellani LW, Kostem E, Furlotte N, Drake TA, Eskin E, Lusis AJ. 2010. A high-resolution association mapping panel for the dissection of complex traits in mice. Genome Res 20(2):281–290. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

