Microfluidic device for rapid digestion of tissues into cellular suspensions
- PMID: 28850139
- PMCID: PMC5614870
- DOI: 10.1039/c7lc00575j
Microfluidic device for rapid digestion of tissues into cellular suspensions
Abstract
The ability to harvest single cells from tissues is currently a bottleneck for cell-based diagnostic technologies, and remains crucial in the fields of tissue engineering and regenerative medicine. Tissues are typically broken down using proteolytic digestion and various mechanical treatments, but success has been limited due to long processing times, low yield, and high manual labor burden. Here, we present a novel microfluidic device that utilizes precision fluid flows to improve the speed and efficiency of tissue digestion. The microfluidic channels were designed to apply hydrodynamic shear forces at discrete locations on tissue specimens up to 1 cm in length and 1 mm in diameter, thereby accelerating digestion through hydrodynamic shear forces and improved enzyme-tissue contact. We show using animal organs that our digestion device with hydro-mincing capabilities was superior to conventional scalpel mincing and digestion based on recovery of DNA and viable single cells. Thus, our microfluidic digestion device can eliminate or reduce the need to mince tissue samples with a scalpel, while reducing sample processing time and preserving cell viability. Another advantage is that downstream microfluidic operations could be integrated to enable advanced cell processing and analysis capabilities. We envision our novel device being used in research and clinical settings to promote single cell-based analysis technologies, as well as to isolate primary, progenitor, and stem cells for use in the fields of tissue engineering and regenerative medicine.
Figures


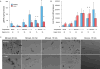

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

