Viral Load Affects the Immune Response to HBV in Mice With Humanized Immune System and Liver
- PMID: 28851562
- PMCID: PMC5733397
- DOI: 10.1053/j.gastro.2017.08.034
Viral Load Affects the Immune Response to HBV in Mice With Humanized Immune System and Liver
Abstract
Background & aims: Hepatitis B virus (HBV) infects hepatocytes, but the mechanisms of the immune response against the virus and how it affects disease progression are unclear.
Methods: We performed studies with BALB/c Rag2-/-Il2rg-/-SirpaNODAlb-uPAtg/tg mice, stably engrafted with human hepatocytes (HUHEP) with or without a human immune system (HIS). HUHEP and HIS-HUHEP mice were given an intraperitoneal injection of HBV. Mononuclear cells were isolated from spleen and liver for analysis by flow cytometry. Liver was analyzed by immunohistochemistry and mRNA levels were measured by quantitative reverse transcription polymerase chain reaction (PCR). Plasma levels of HBV DNA were quantified by PCR reaction, and antigen-specific antibodies were detected by immunocytochemistry of HBV-transfected BHK-21 cells.
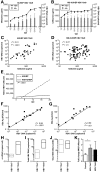
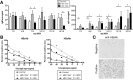
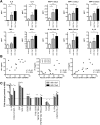
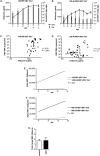
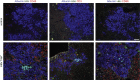
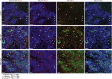
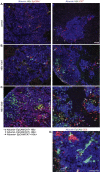
Results: Following HBV infection, a complete viral life cycle, with production of HBV DNA, hepatitis B e (HBe), core (HBc) and surface (HBs) antigens, and covalently closed circular DNA, was observed in HUHEP and HIS-HUHEP mice. HBV replicated unrestricted in HUHEP mice resulting in high viral titers without pathologic effects. In contrast, HBV-infected HIS-HUHEP mice developed chronic hepatitis with 10-fold lower titers and antigen-specific IgGs, (anti-HBs, anti-HBc), consistent with partial immune control. HBV-infected HIS-HUHEP livers contained infiltrating Kupffer cells, mature activated natural killer cells (CD69+), and PD-1+ effector memory T cells (CD45RO+). Reducing the viral inoculum resulted in more efficient immune control. Plasma from HBV-infected HIS-HUHEP mice had increased levels of inflammatory and immune-suppressive cytokines (C-X-C motif chemokine ligand 10 and interleukin 10), which correlated with populations of intrahepatic CD4+ T cells (CD45RO+PD-1+). Mice with high levels of viremia had HBV-infected liver progenitor cells. Giving the mice the nucleoside analogue entecavir reduced viral loads and decreased liver inflammation.
Conclusion: In HIS-HUHEP mice, HBV infection completes a full life cycle and recapitulates some of the immunopathology observed in patients with chronic infection. Inoculation with different viral loads led to different immune responses and levels of virus control. We found HBV to infect liver progenitor cells, which could be involved in hepatocellular carcinogenesis. This is an important new system to study anti-HBV immune responses and screen for combination therapies against hepatotropic viruses.
Keywords: CXCL10; HBe; IL10; Mouse Model; NK Cell.
Copyright © 2017. Published by Elsevier Inc.
Figures







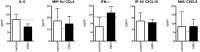
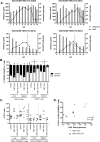
References
-
- Zeisel M.B., Lucifora J., Mason W.S. Towards an HBV cure: state-of-the-art and unresolved questions-report of the ANRS workshop on HBV cure. Gut. 2015;64:1314–1326. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

