Genome-Wide Spectra of Transcription Insertions and Deletions Reveal That Slippage Depends on RNA:DNA Hybrid Complementarity
- PMID: 28851848
- PMCID: PMC5574713
- DOI: 10.1128/mBio.01230-17
Genome-Wide Spectra of Transcription Insertions and Deletions Reveal That Slippage Depends on RNA:DNA Hybrid Complementarity
Abstract
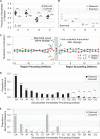
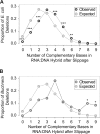
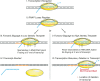
Advances in sequencing technologies have enabled direct quantification of genome-wide errors that occur during RNA transcription. These errors occur at rates that are orders of magnitude higher than rates during DNA replication, but due to technical difficulties such measurements have been limited to single-base substitutions and have not yet quantified the scope of transcription insertions and deletions. Previous reporter gene assay findings suggested that transcription indels are produced exclusively by elongation complex slippage at homopolymeric runs, so we enumerated indels across the protein-coding transcriptomes of Escherichia coli and Buchnera aphidicola, which differ widely in their genomic base compositions and incidence of repeat regions. As anticipated from prior assays, transcription insertions prevailed in homopolymeric runs of A and T; however, transcription deletions arose in much more complex sequences and were rarely associated with homopolymeric runs. By reconstructing the relocated positions of the elongation complex as inferred from the sequences inserted or deleted during transcription, we show that continuation of transcription after slippage hinges on the degree of nucleotide complementarity within the RNA:DNA hybrid at the new DNA template location.IMPORTANCE The high level of mistakes generated during transcription can result in the accumulation of malfunctioning and misfolded proteins which can alter global gene regulation and in the expenditure of energy to degrade these nonfunctional proteins. The transcriptome-wide occurrence of base substitutions has been elucidated in bacteria, but information on transcription insertions and deletions-errors that potentially have more dire effects on protein function-is limited to reporter gene constructs. Here, we capture the transcriptome-wide spectrum of insertions and deletions in Escherichia coli and Buchnera aphidicola and show that they occur at rates approaching those of base substitutions. Knowledge of the full extent of sequences subject to transcription indels supports a new model of bacterial transcription slippage, one that relies on the number of complementary bases between the transcript and the DNA template to which it slipped.
Keywords: Buchnera; Escherichia coli; RNA polymerase; elongation complex; error rates; mutation; transcription; transcription slippage.
Copyright © 2017 Traverse and Ochman.
Figures


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
