Engineering of versatile redox partner fusions that support monooxygenase activity of functionally diverse cytochrome P450s
- PMID: 28852040
- PMCID: PMC5575160
- DOI: 10.1038/s41598-017-10075-w
Engineering of versatile redox partner fusions that support monooxygenase activity of functionally diverse cytochrome P450s
Abstract
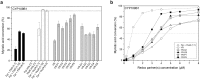
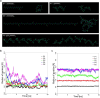
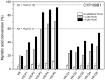
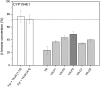
Most bacterial cytochrome P450 monooxygenases (P450s or CYPs) require two redox partner proteins for activity. To reduce complexity of the redox chain, the Bacillus subtilis flavodoxin YkuN (Y) was fused to the Escherichia coli flavodoxin reductase Fpr (R), and activity was tuned by placing flexible (GGGGS)n or rigid ([E/L]PPPP)n linkers (n = 1-5) in between. P-linker constructs typically outperformed their G-linker counterparts, with superior performance of YR-P5, which carries linker ([E/L]PPPP)5. Molecular dynamics simulations demonstrated that ([E/L]PPPP)n linkers are intrinsically rigid, whereas (GGGGS)n linkers are highly flexible and biochemical experiments suggest a higher degree of separation between the fusion partners in case of long rigid P-linkers. The catalytic properties of the individual redox partners were best preserved in the YR-P5 construct. In comparison to the separate redox partners, YR-P5 exhibited attenuated rates of NADPH oxidation and heme iron (III) reduction, while coupling efficiency was improved (28% vs. 49% coupling with B. subtilis CYP109B1, and 44% vs. 50% with Thermobifida fusca CYP154E1). In addition, YR-P5 supported monooxygenase activity of the CYP106A2 from Bacillus megaterium and bovine CYP21A2. The versatile YR-P5 may serve as a non-physiological electron transfer system for exploitation of the catalytic potential of other P450s.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
Design and improvement of artificial redox modules by molecular fusion of flavodoxin and flavodoxin reductase from Escherichia coli.Sci Rep. 2015 Jul 16;5:12158. doi: 10.1038/srep12158. Sci Rep. 2015. PMID: 26177696 Free PMC article.
-
Expression, purification, and characterization of Bacillus subtilis cytochromes P450 CYP102A2 and CYP102A3: flavocytochrome homologues of P450 BM3 from Bacillus megaterium.Biochemistry. 2004 May 11;43(18):5474-87. doi: 10.1021/bi035904m. Biochemistry. 2004. PMID: 15122913
-
Cytochrome P450--redox partner fusion enzymes.Biochim Biophys Acta. 2007 Mar;1770(3):345-59. doi: 10.1016/j.bbagen.2006.08.018. Epub 2006 Aug 30. Biochim Biophys Acta. 2007. PMID: 17023115 Review.
-
Expression and characterization of the two flavodoxin proteins of Bacillus subtilis, YkuN and YkuP: biophysical properties and interactions with cytochrome P450 BioI.Biochemistry. 2004 Oct 5;43(39):12390-409. doi: 10.1021/bi049131t. Biochemistry. 2004. PMID: 15449930
-
Biological diversity of cytochrome P450 redox partner systems.Adv Exp Med Biol. 2015;851:299-317. doi: 10.1007/978-3-319-16009-2_11. Adv Exp Med Biol. 2015. PMID: 26002740 Review.
Cited by
-
Engineering Electron Transfer Pathway of Cytochrome P450s.Molecules. 2024 May 24;29(11):2480. doi: 10.3390/molecules29112480. Molecules. 2024. PMID: 38893355 Free PMC article. Review.
-
Genetic fusion of P450 BM3 and formate dehydrogenase towards self-sufficient biocatalysts with enhanced activity.Sci Rep. 2021 Nov 4;11(1):21706. doi: 10.1038/s41598-021-00957-5. Sci Rep. 2021. PMID: 34737365 Free PMC article.
-
Multi-step pathway engineering in probiotic Saccharomyces boulardii for abscisic acid production in the gut.Metab Eng Commun. 2025 May 30;20:e00263. doi: 10.1016/j.mec.2025.e00263. eCollection 2025 Jun. Metab Eng Commun. 2025. PMID: 40530243 Free PMC article.
-
Regio- and Stereoselective Steroid Hydroxylation at C7 by Cytochrome P450 Monooxygenase Mutants.Angew Chem Int Ed Engl. 2020 Jul 20;59(30):12499-12505. doi: 10.1002/anie.202003139. Epub 2020 May 25. Angew Chem Int Ed Engl. 2020. PMID: 32243054 Free PMC article.
-
Heterologous caffeic acid biosynthesis in Escherichia coli is affected by choice of tyrosine ammonia lyase and redox partners for bacterial Cytochrome P450.Microb Cell Fact. 2020 Feb 11;19(1):26. doi: 10.1186/s12934-020-01300-9. Microb Cell Fact. 2020. PMID: 32046741 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

