Metabolic profiling of triple-negative breast cancer cells reveals metabolic vulnerabilities
- PMID: 28852500
- PMCID: PMC5568171
- DOI: 10.1186/s40170-017-0168-x
Metabolic profiling of triple-negative breast cancer cells reveals metabolic vulnerabilities
Abstract
Background: Among breast cancers, the triple-negative breast cancer (TNBC) subtype has the worst prognosis with no approved targeted therapies and only standard chemotherapy as the backbone of systemic therapy. Unique metabolic changes in cancer progression provide innovative therapeutic opportunities. The receptor tyrosine kinases (RTKs) epidermal growth factor receptor (EGFR), and MET receptor are highly expressed in TNBC, making both promising therapeutic targets. RTK signaling profoundly alters cellular metabolism by increasing glucose consumption and subsequently diverting glucose carbon sources into metabolic pathways necessary to support the tumorigenesis. Therefore, detailed metabolic profiles of TNBC subtypes and their response to tyrosine kinase inhibitors may identify therapeutic sensitivities.
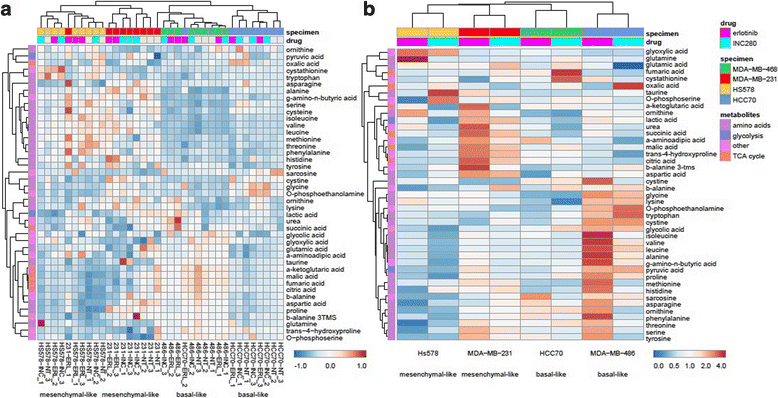
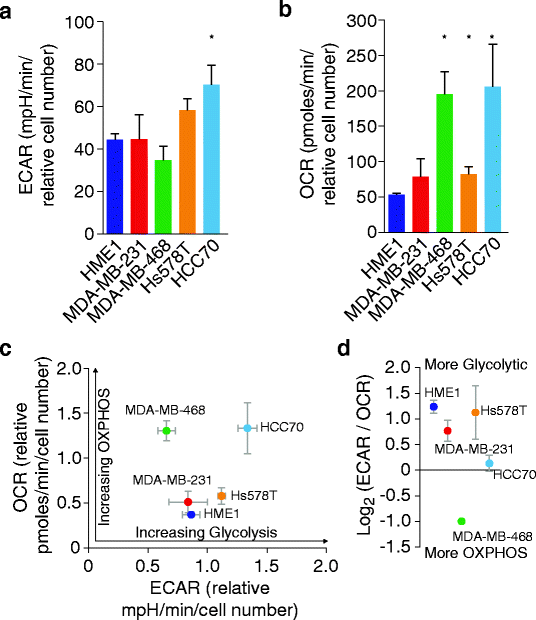
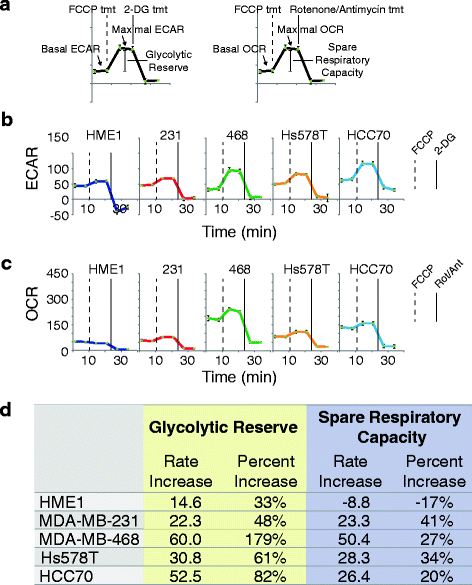
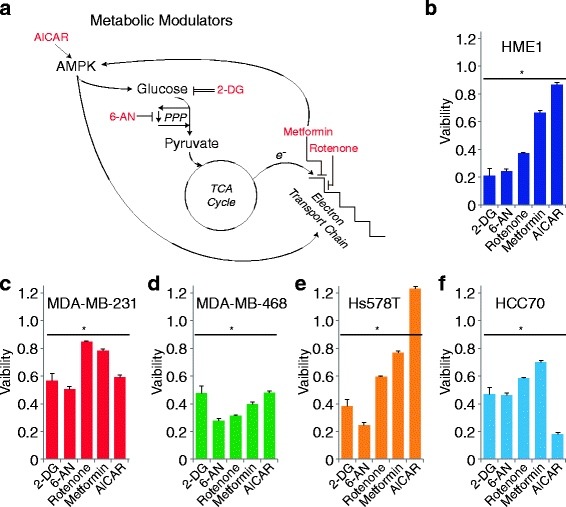
Methods: We quantified the metabolic profiles of TNBC cell lines representing multiple TNBC subtypes using gas chromatography mass spectrometry. In addition, we subjected MDA-MB-231, MDA-MB-468, Hs578T, and HCC70 cell lines to metabolic flux analysis of basal and maximal glycolytic and mitochondrial oxidative rates. Metabolic pool size and flux measurements were performed in the presence and absence of the MET inhibitor, INC280/capmatinib, and the EGFR inhibitor, erlotinib. Further, the sensitivities of these cells to modulators of core metabolic pathways were determined. In addition, we annotated a rate-limiting metabolic enzymes library and performed a siRNA screen in combination with MET or EGFR inhibitors to validate synergistic effects.
Results: TNBC cell line models displayed significant metabolic heterogeneity with respect to basal and maximal metabolic rates and responses to RTK and metabolic pathway inhibitors. Comprehensive systems biology analysis of metabolic perturbations, combined siRNA and tyrosine kinase inhibitor screens identified a core set of TCA cycle and fatty acid pathways whose perturbation sensitizes TNBC cells to small molecule targeting of receptor tyrosine kinases.
Conclusions: Similar to the genomic heterogeneity observed in TNBC, our results reveal metabolic heterogeneity among TNBC subtypes and demonstrate that understanding metabolic profiles and drug responses may prove valuable in targeting TNBC subtypes and identifying therapeutic susceptibilities in TNBC patients. Perturbation of metabolic pathways sensitizes TNBC to inhibition of receptor tyrosine kinases. Such metabolic vulnerabilities offer promise for effective therapeutic targeting for TNBC patients.
Keywords: Metabolic inhibitor; Metabolism; Rate-limiting enzymes; Receptor tyrosine kinase; Triple-negative breast cancer; Tyrosine kinase inhibitor.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
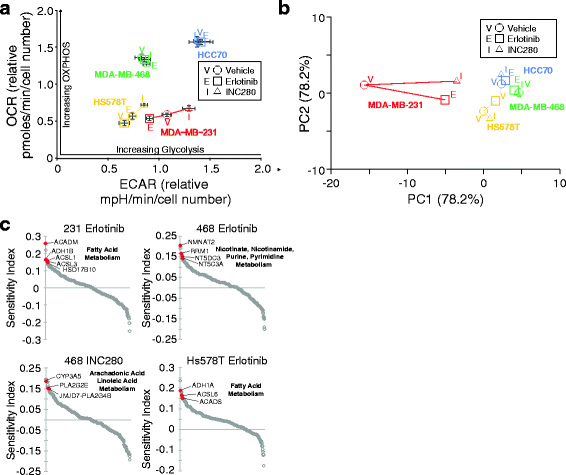
References
-
- Liedtke C, Mazouni C, Hess KR, André F, Tordai A, Mejia JA, Symmans WF, Gonzalez-Angulo AM, Hennessy B, Green M, Cristofanilli M, Hortobagyi GN, Pusztai L. Response to neoadjuvant therapy and long-term survival in patients with triple-negative breast cancer. J Clin Oncol. 2008;26:1275–1281. doi: 10.1200/JCO.2007.14.4147. - DOI - PubMed
-
- Burstein MD, Tsimelzon A, Poage GM, Covington KR, Contreras A, Fuqua SAW, Savage MI, Osborne CK, Hilsenbeck SG, Chang JC, Mills GB, Lau CC, Brown PH. Comprehensive genomic analysis identifies novel subtypes and targets of triple-negative breast cancer. Clin Cancer Res. 2015;21:1688–1698. doi: 10.1158/1078-0432.CCR-14-0432. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

