Further investigations of the W-test for pairwise epistasis testing
- PMID: 28852712
- PMCID: PMC5553086
- DOI: 10.12688/wellcomeopenres.11926.1
Further investigations of the W-test for pairwise epistasis testing
Abstract
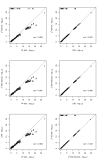
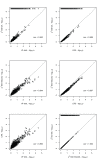
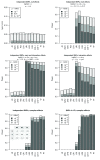
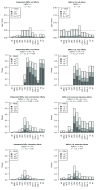
Background: In a recent paper, a novel W-test for pairwise epistasis testing was proposed that appeared, in computer simulations, to have higher power than competing alternatives. Application to genome-wide bipolar data detected significant epistasis between SNPs in genes of relevant biological function. Network analysis indicated that the implicated genes formed two separate interaction networks, each containing genes highly related to autism and neurodegenerative disorders. Methods: Here we investigate further the properties and performance of the W-test via theoretical evaluation, computer simulations and application to real data. Results: We demonstrate that, for common variants, the W-test is closely related to several existing tests of association allowing for interaction, including logistic regression on 8 degrees of freedom, although logistic regression can show inflated type I error for low minor allele frequencies, whereas the W-test shows good/conservative type I error control. Although in some situations the W-test can show higher power, logistic regression is not limited to tests on 8 degrees of freedom but can instead be tailored to impose greater structure on the assumed alternative hypothesis, offering a power advantage when the imposed structure matches the true structure. Conclusions: The W-test is a potentially useful method for testing for association - without necessarily implying interaction - between genetic variants disease, particularly when one or more of the genetic variants are rare. For common variants, the advantages of the W-test are less clear, and, indeed, there are situations where existing methods perform better. In our investigations, we further uncover a number of problems with the practical implementation and application of the W-test (to bipolar disorder) previously described, apparently due to inadequate use of standard data quality-control procedures. This observation leads us to urge caution in interpretation of the previously-presented results, most of which we consider are highly likely to be artefacts.
Keywords: GWAS; Interactions; contingency table; epistasis; quality control.
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures
Similar articles
-
A fast and powerful W-test for pairwise epistasis testing.Nucleic Acids Res. 2016 Jul 8;44(12):e115. doi: 10.1093/nar/gkw347. Epub 2016 Apr 25. Nucleic Acids Res. 2016. PMID: 27112568 Free PMC article.
-
Improved statistics for genome-wide interaction analysis.PLoS Genet. 2012;8(4):e1002625. doi: 10.1371/journal.pgen.1002625. Epub 2012 Apr 5. PLoS Genet. 2012. PMID: 22496670 Free PMC article.
-
Performance of epistasis detection methods in semi-simulated GWAS.BMC Bioinformatics. 2018 Jun 18;19(1):231. doi: 10.1186/s12859-018-2229-8. BMC Bioinformatics. 2018. PMID: 29914375 Free PMC article.
-
Folic acid supplementation and malaria susceptibility and severity among people taking antifolate antimalarial drugs in endemic areas.Cochrane Database Syst Rev. 2022 Feb 1;2(2022):CD014217. doi: 10.1002/14651858.CD014217. Cochrane Database Syst Rev. 2022. PMID: 36321557 Free PMC article.
-
Impact of summer programmes on the outcomes of disadvantaged or 'at risk' young people: A systematic review.Campbell Syst Rev. 2024 Jun 13;20(2):e1406. doi: 10.1002/cl2.1406. eCollection 2024 Jun. Campbell Syst Rev. 2024. PMID: 38873396 Free PMC article. Review.
References
-
- Newman SC: Biostatistical Methods in Epidemiology. Wiley,2001. 10.1002/0471272612 - DOI
-
- Agresti A: Categorical Data Analysis, 3rd Edition. Wiley,2013. Reference Source
-
- Pearson K: On the criterion that a given system of deviations from the probable in the case of a correlated system of variables is such that it can be reasonably supposed to have arisen from random sampling. Philosophical Magazine Series 5. 1900;50(302):157–175. 10.1080/14786440009463897 - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

