Identification of metal ion binding sites based on amino acid sequences
- PMID: 28854211
- PMCID: PMC5576659
- DOI: 10.1371/journal.pone.0183756
Identification of metal ion binding sites based on amino acid sequences
Abstract
The identification of metal ion binding sites is important for protein function annotation and the design of new drug molecules. This study presents an effective method of analyzing and identifying the binding residues of metal ions based solely on sequence information. Ten metal ions were extracted from the BioLip database: Zn2+, Cu2+, Fe2+, Fe3+, Ca2+, Mg2+, Mn2+, Na+, K+ and Co2+. The analysis showed that Zn2+, Cu2+, Fe2+, Fe3+, and Co2+ were sensitive to the conservation of amino acids at binding sites, and promising results can be achieved using the Position Weight Scoring Matrix algorithm, with an accuracy of over 79.9% and a Matthews correlation coefficient of over 0.6. The binding sites of other metals can also be accurately identified using the Support Vector Machine algorithm with multifeature parameters as input. In addition, we found that Ca2+ was insensitive to hydrophobicity and hydrophilicity information and Mn2+ was insensitive to polarization charge information. An online server was constructed based on the framework of the proposed method and is freely available at http://60.31.198.140:8081/metal/HomePage/HomePage.html.
Conflict of interest statement
Figures
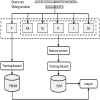
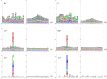
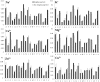

References
-
- Ibers J A, Holm R H. Modeling coordination sites in metallobiomolecules. Science, 1980, 209(4453):223–35. - PubMed
-
- Tainer J A, Roberts V A, Getzoff E D. Metal-binding sites in proteins. Current Opinion in Biotechnology, 1991, 2(4):582–91. - PubMed
-
- Degtyarenko K. Bioinorganic motifs: towards functional classification of metalloproteins. Bioinformatics, 2000, 16(10):851–64. - PubMed
-
- Reif D W. Ferritin as a source of iron for oxidative damage. Free Radical Biology & Medicine, 1992, 12(5):417–427. - PubMed
-
- Tupler R, Perini G, Green M R. Expressing the human genome. Nature, 2001, 409(409):832–3. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

