CgII cleaves DNA using a mechanism distinct from other ATP-dependent restriction endonucleases
- PMID: 28854738
- PMCID: PMC5737866
- DOI: 10.1093/nar/gkx580
CgII cleaves DNA using a mechanism distinct from other ATP-dependent restriction endonucleases
Abstract
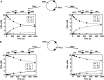
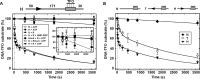
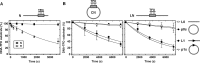
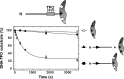
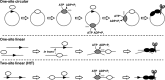
The restriction endonuclease CglI from Corynebacterium glutamicum recognizes an asymmetric 5'-GCCGC-3' site and cleaves the DNA 7 and 6/7 nucleotides downstream on the top and bottom DNA strands, respectively, in an NTP-hydrolysis dependent reaction. CglI is composed of two different proteins: an endonuclease (R.CglI) and a DEAD-family helicase-like ATPase (H.CglI). These subunits form a heterotetrameric complex with R2H2 stoichiometry. However, the R2H2·CglI complex has only one nuclease active site sufficient to cut one DNA strand suggesting that two complexes are required to introduce a double strand break. Here, we report studies to evaluate the DNA cleavage mechanism of CglI. Using one- and two-site circular DNA substrates we show that CglI does not require two sites on the same DNA for optimal catalytic activity. However, one-site linear DNA is a poor substrate, supporting a mechanism where CglI complexes must communicate along the one-dimensional DNA contour before cleavage is activated. Based on experimental data, we propose that adenosine triphosphate (ATP) hydrolysis by CglI produces translocation on DNA preferentially in a downstream direction from the target, although upstream translocation is also possible. Our results are consistent with a mechanism of CglI action that is distinct from that of other ATP-dependent restriction-modification enzymes.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



Similar articles
-
The H-subunit of the restriction endonuclease CglI contains a prototype DEAD-Z1 helicase-like motor.Nucleic Acids Res. 2018 Mar 16;46(5):2560-2572. doi: 10.1093/nar/gky107. Nucleic Acids Res. 2018. PMID: 29471489 Free PMC article.
-
DNA cleavage by CgII and NgoAVII requires interaction between N- and R-proteins and extensive nucleotide hydrolysis.Nucleic Acids Res. 2014 Dec 16;42(22):13887-96. doi: 10.1093/nar/gku1236. Epub 2014 Nov 27. Nucleic Acids Res. 2014. PMID: 25429977 Free PMC article.
-
Complex restriction enzymes: NTP-driven molecular motors.Biochimie. 2002 Nov;84(11):1047-59. doi: 10.1016/s0300-9084(02)00020-2. Biochimie. 2002. PMID: 12595133 Review.
-
Subunit assembly and mode of DNA cleavage of the type III restriction endonucleases EcoP1I and EcoP15I.J Mol Biol. 2001 Feb 23;306(3):417-31. doi: 10.1006/jmbi.2000.4411. J Mol Biol. 2001. PMID: 11178902
-
The mechanism of ATP-dependent RNA unwinding by DEAD box proteins.Biol Chem. 2009 Dec;390(12):1237-50. doi: 10.1515/BC.2009.135. Biol Chem. 2009. PMID: 19747077 Review.
Cited by
-
The H-subunit of the restriction endonuclease CglI contains a prototype DEAD-Z1 helicase-like motor.Nucleic Acids Res. 2018 Mar 16;46(5):2560-2572. doi: 10.1093/nar/gky107. Nucleic Acids Res. 2018. PMID: 29471489 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

