Function and Safety of Lentivirus-Mediated Gene Transfer for CSF2RA-Deficiency
- PMID: 28854814
- PMCID: PMC5734162
- DOI: 10.1089/hgtb.2017.092
Function and Safety of Lentivirus-Mediated Gene Transfer for CSF2RA-Deficiency
Abstract
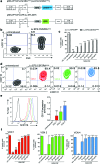
Hereditary pulmonary alveolar proteinosis (hPAP) is a rare disorder of pulmonary surfactant accumulation and hypoxemic respiratory failure caused by mutations in CSF2RA (encoding the granulocyte/macrophage colony-stimulating factor [GM-CSF] receptor α-chain [CD116]), which results in reduced GM-CSF-dependent pulmonary surfactant clearance by alveolar macrophages. While no pharmacologic therapy currently exists for hPAP, it was recently demonstrated that endotracheal instillation of wild-type or gene-corrected mononuclear phagocytes (pulmonary macrophage transplantation [PMT]) results in a significant and durable therapeutic efficacy in a validated murine model of hPAP. To facilitate the translation of PMT therapy to human hPAP patients, a self-inactivating (SIN) lentiviral vector was generated expressing a codon-optimized human CSF2RA-cDNA driven from an EF1α short promoter (Lv.EFS.CSF2RAcoop), and a series of nonclinical efficacy and safety studies were performed in cultured macrophage cell lines and primary human cells. Studies in cytokine-dependent Ba/F3 cells demonstrated efficient transduction, vector-derived CD116 expression proportional to vector copy number, and GM-CSF-dependent cell survival and proliferation. Using a novel cell line constructed to express a normal GM-CSF receptor β subunit and a dysfunctional α subunit (due to a function-altering CSF2RAG196R mutation) that reflects the macrophage disease phenotype of hPAP patients, it was demonstrated that Lv.EFS.CSF2RAcoop transduction restored GM-CSF receptor function. Further, Lv.EFS.CSF2RAcoop transduction of healthy primary CD34+ cells did not adversely affect cell proliferation or affect the cell differentiation program. Results demonstrate Lv.EFS.CSF2RAcoop reconstituted GM-CSF receptor α expression, restoring GM-CSF signaling in hPAP macrophages, and had no adverse effects in the intended target cells, thus supporting testing of PMT therapy of hPAP in humans.
Keywords: CSF2RA; HSCs; hPAP; hematopoietic gene therapy; lentivirus; macrophages.
Conflict of interest statement
The authors have no commercial, proprietary, or financial interest in the products or companies described in this article.
Figures



Similar articles
-
Long-Term Safety and Efficacy of Gene-Pulmonary Macrophage Transplantation Therapy of PAP in Csf2ra-/- Mice.Mol Ther. 2019 Sep 4;27(9):1597-1611. doi: 10.1016/j.ymthe.2019.06.010. Epub 2019 Jul 2. Mol Ther. 2019. PMID: 31326401 Free PMC article.
-
A murine model of hereditary pulmonary alveolar proteinosis caused by homozygous Csf2ra gene disruption.Am J Physiol Lung Cell Mol Physiol. 2022 Mar 1;322(3):L438-L448. doi: 10.1152/ajplung.00175.2021. Epub 2022 Jan 19. Am J Physiol Lung Cell Mol Physiol. 2022. PMID: 35043685 Free PMC article.
-
Gene correction of human induced pluripotent stem cells repairs the cellular phenotype in pulmonary alveolar proteinosis.Am J Respir Crit Care Med. 2014 Jan 15;189(2):167-82. doi: 10.1164/rccm.201306-1012OC. Am J Respir Crit Care Med. 2014. PMID: 24279725
-
Immune dysregulation in the pathogenesis of pulmonary alveolar proteinosis.Curr Allergy Asthma Rep. 2010 Sep;10(5):320-5. doi: 10.1007/s11882-010-0134-y. Curr Allergy Asthma Rep. 2010. PMID: 20623372 Review.
-
Pulmonary alveolar proteinosis, a primary immunodeficiency of impaired GM-CSF stimulation of macrophages.Curr Opin Immunol. 2009 Oct;21(5):514-21. doi: 10.1016/j.coi.2009.09.004. Epub 2009 Sep 30. Curr Opin Immunol. 2009. PMID: 19796925 Free PMC article. Review.
Cited by
-
Emerging Treatments for Childhood Interstitial Lung Disease.Paediatr Drugs. 2024 Jan;26(1):19-30. doi: 10.1007/s40272-023-00603-9. Epub 2023 Nov 10. Paediatr Drugs. 2024. PMID: 37948041 Free PMC article.
-
Ex Vivo Generation of CAR Macrophages from Hematopoietic Stem and Progenitor Cells for Use in Cancer Therapy.Cells. 2022 Mar 15;11(6):994. doi: 10.3390/cells11060994. Cells. 2022. PMID: 35326445 Free PMC article.
-
Pulmonary Alveolar Proteinosis and new therapeutic concepts.Klin Padiatr. 2024 Feb;236(2):73-79. doi: 10.1055/a-2233-1243. Epub 2024 Jan 29. Klin Padiatr. 2024. PMID: 38286410 Free PMC article.
-
Effective hematopoietic stem cell-based gene therapy in a murine model of hereditary pulmonary alveolar proteinosis.Haematologica. 2020 Apr;105(4):1147-1157. doi: 10.3324/haematol.2018.214866. Epub 2019 Jul 9. Haematologica. 2020. PMID: 31289207 Free PMC article.
-
New approaches to moderate CRISPR-Cas9 activity: Addressing issues of cellular uptake and endosomal escape.Mol Ther. 2022 Jan 5;30(1):32-46. doi: 10.1016/j.ymthe.2021.06.003. Epub 2021 Jun 4. Mol Ther. 2022. PMID: 34091053 Free PMC article. Review.
References
-
- Shi Y, Liu CH, Roberts AI, et al. . Granulocyte-macrophage colony-stimulating factor (GM-CSF) and T-cell responses: what we do and don't know. Cell Res 2006;16:126–133 - PubMed
-
- Nishinakamura R, Miyajima A, Mee PJ, et al. . Hematopoiesis in mice lacking the entire granulocyte-macrophage colony-stimulating factor/interleukin-3/interleukin-5 functions. Blood 1996;88:2458–2464 - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

