Protein complex formation during denitrification by Pseudomonas aeruginosa
- PMID: 28857512
- PMCID: PMC5658584
- DOI: 10.1111/1751-7915.12851
Protein complex formation during denitrification by Pseudomonas aeruginosa
Erratum in
-
Corrigendum.Microb Biotechnol. 2018 Mar;11(2):431. doi: 10.1111/1751-7915.13048. Microb Biotechnol. 2018. PMID: 29442450 Free PMC article. No abstract available.
Abstract
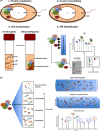
The most efficient means of generating cellular energy is through aerobic respiration. Under anaerobic conditions, several prokaryotes can replace oxygen by nitrate as final electron acceptor. During denitrification, nitrate is reduced via nitrite, NO and N2 O to molecular nitrogen (N2 ) by four membrane-localized reductases with the simultaneous formation of an ion gradient for ATP synthesis. These four multisubunit enzyme complexes are coupled in four electron transport chains to electron donating primary dehydrogenases and intermediate electron transfer proteins. Many components require membrane transport and insertion, complex assembly and cofactor incorporation. All these processes are mediated by fine-tuned stable and transient protein-protein interactions. Recently, an interactomic approach was used to determine the exact protein-protein interactions involved in the assembly of the denitrification apparatus of Pseudomonas aeruginosa. Both subunits of the NO reductase NorBC, combined with the flavoprotein NosR, serve as a membrane-localized assembly platform for the attachment of the nitrate reductase NarGHI, the periplasmic nitrite reductase NirS via its maturation factor NirF and the N2 O reductase NosZ through NosR. A nitrate transporter (NarK2), the corresponding regulatory system NarXL, various nitrite (NirEJMNQ) and N2 O reductase (NosFL) maturation proteins are also part of the complex. Primary dehydrogenases, ATP synthase, most enzymes of the TCA cycle, and the SEC protein export system, as well as a number of other proteins, were found to interact with the denitrification complex. Finally, a protein complex composed of the flagella protein FliC, nitrite reductase NirS and the chaperone DnaK required for flagella formation was found in the periplasm of P. aeruginosa. This work demonstrated that the interactomic approach allows for the identification and characterization of stable and transient protein-protein complexes and interactions involved in the assembly and function of multi-enzyme complexes.
© 2017 The Authors. Microbial Biotechnology published by John Wiley & Sons Ltd and Society for Applied Microbiology.
Figures


Similar articles
-
Protein Network of the Pseudomonas aeruginosa Denitrification Apparatus.J Bacteriol. 2016 Apr 14;198(9):1401-13. doi: 10.1128/JB.00055-16. Print 2016 May. J Bacteriol. 2016. PMID: 26903416 Free PMC article.
-
A Periplasmic Complex of the Nitrite Reductase NirS, the Chaperone DnaK, and the Flagellum Protein FliC Is Essential for Flagellum Assembly and Motility in Pseudomonas aeruginosa.J Bacteriol. 2015 Oct;197(19):3066-75. doi: 10.1128/JB.00415-15. Epub 2015 Jul 13. J Bacteriol. 2015. PMID: 26170416 Free PMC article.
-
Localization of denitrification genes on the chromosomal map of Pseudomonas aeruginosa.Microbiology (Reading). 1998 Feb;144 ( Pt 2):441-448. doi: 10.1099/00221287-144-2-441. Microbiology (Reading). 1998. PMID: 9493381
-
Metabolic regulation including anaerobic metabolism in Paracoccus denitrificans.J Bioenerg Biomembr. 1991 Apr;23(2):163-85. doi: 10.1007/BF00762216. J Bioenerg Biomembr. 1991. PMID: 2050653 Review.
-
Denitrification and nitrite reduction: Pseudomonas aeruginosa nitrite-reductase.Biochimie. 1984 Apr;66(4):259-89. doi: 10.1016/0300-9084(84)90005-1. Biochimie. 1984. PMID: 6331530 Review.
Cited by
-
The Potential for Redox-Active Metabolites To Enhance or Unlock Anaerobic Survival Metabolisms in Aerobes.J Bacteriol. 2020 May 11;202(11):e00797-19. doi: 10.1128/JB.00797-19. Print 2020 May 11. J Bacteriol. 2020. PMID: 32071098 Free PMC article. Review.
-
Impact of dissolved oxygen and loading rate on NH3 oxidation and N2 production mechanisms in activated sludge treatment of sewage.Microb Biotechnol. 2021 Mar;14(2):419-429. doi: 10.1111/1751-7915.13599. Epub 2020 Jun 2. Microb Biotechnol. 2021. PMID: 32488999 Free PMC article.
-
Mechanisms controlling bacterial infection in myeloid cells under hypoxic conditions.Cell Mol Life Sci. 2021 Mar;78(5):1887-1907. doi: 10.1007/s00018-020-03684-8. Epub 2020 Oct 30. Cell Mol Life Sci. 2021. PMID: 33125509 Free PMC article. Review.
-
Nitrogen Removal Performance and Metabolic Pathways Analysis of a Novel Aerobic Denitrifying Halotolerant Pseudomonas balearica strain RAD-17.Microorganisms. 2020 Jan 2;8(1):72. doi: 10.3390/microorganisms8010072. Microorganisms. 2020. PMID: 31906569 Free PMC article.
-
The O2-independent pathway of ubiquinone biosynthesis is essential for denitrification in Pseudomonas aeruginosa.J Biol Chem. 2020 Jul 3;295(27):9021-9032. doi: 10.1074/jbc.RA120.013748. Epub 2020 May 14. J Biol Chem. 2020. PMID: 32409583 Free PMC article.
References
-
- Acin‐Perez, R. , and Enriquez, J.A. (2014) The function of the respiratory supercomplexes: the plasticity model. Biochim Biophys Acta 1837: 444–450. - PubMed
-
- Al Shweiki, M.R. , Monchgesang, S. , Majovsky, P. , Thieme, D. , Trutschel, D. , and Hoehenwarter, W. (2017) Assessment of label‐free quantification in discovery proteomics and impact of technological factors and natural variability of protein abundance. J Proteome Res 16: 1410–1424. - PubMed
-
- Alcazar‐Fabra, M. , Navas, P. , and Brea‐Calvo, G. (2016) Coenzyme Q biosynthesis and its role in the respiratory chain structure. Biochim Biophys Acta 1857: 1073–1078. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

