Protein-Protein Interactions in Virus-Host Systems
- PMID: 28861068
- PMCID: PMC5562681
- DOI: 10.3389/fmicb.2017.01557
Protein-Protein Interactions in Virus-Host Systems
Abstract
To study virus-host protein interactions, knowledge about viral and host protein architectures and repertoires, their particular evolutionary mechanisms, and information on relevant sources of biological data is essential. The purpose of this review article is to provide a thorough overview about these aspects. Protein domains are basic units defining protein interactions, and the uniqueness of viral domain repertoires, their mode of evolution, and their roles during viral infection make viruses interesting models of study. Mutations at protein interfaces can reduce or increase their binding affinities by changing protein electrostatics and structural properties. During the course of a viral infection, both pathogen and cellular proteins are constantly competing for binding partners. Endogenous interfaces mediating intraspecific interactions-viral-viral or host-host interactions-are constantly targeted and inhibited by exogenous interfaces mediating viral-host interactions. From a biomedical perspective, blocking such interactions is the main mechanism underlying antiviral therapies. Some proteins are able to bind multiple partners, and their modes of interaction define how fast these "hub proteins" evolve. "Party hubs" have multiple interfaces; they establish simultaneous/stable (domain-domain) interactions, and tend to evolve slowly. On the other hand, "date hubs" have few interfaces; they establish transient/weak (domain-motif) interactions by means of short linear peptides (15 or fewer residues), and can evolve faster. Viral infections are mediated by several protein-protein interactions (PPIs), which can be represented as networks (protein interaction networks, PINs), with proteins being depicted as nodes, and their interactions as edges. It has been suggested that viral proteins tend to establish interactions with more central and highly connected host proteins. In an evolutionary arms race, viral and host proteins are constantly changing their interface residues, either to evade or to optimize their binding capabilities. Apart from gaining and losing interactions via rewiring mechanisms, virus-host PINs also evolve via gene duplication (paralogy); conservation (orthology); horizontal gene transfer (HGT) (xenology); and molecular mimicry (convergence). The last sections of this review focus on PPI experimental approaches and their limitations, and provide an overview of sources of biomolecular data for studying virus-host protein interactions.
Keywords: PPI; databases; integrative biology; molecular evolution; protein interaction networks; structural biology; viral evolution; virus-host interactions.
Figures

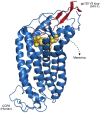




References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

