Comparative Genomic Analysis of Neutrophilic Iron(II) Oxidizer Genomes for Candidate Genes in Extracellular Electron Transfer
- PMID: 28871245
- PMCID: PMC5566968
- DOI: 10.3389/fmicb.2017.01584
Comparative Genomic Analysis of Neutrophilic Iron(II) Oxidizer Genomes for Candidate Genes in Extracellular Electron Transfer
Abstract
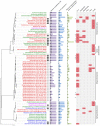
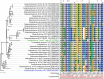
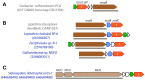
Extracellular electron transfer (EET) is recognized as a key biochemical process in circumneutral pH Fe(II)-oxidizing bacteria (FeOB). In this study, we searched for candidate EET genes in 73 neutrophilic FeOB genomes, among which 43 genomes are complete or close-to-complete and the rest have estimated genome completeness ranging from 5 to 91%. These neutrophilic FeOB span members of the microaerophilic, anaerobic phototrophic, and anaerobic nitrate-reducing FeOB groups. We found that many microaerophilic and several anaerobic FeOB possess homologs of Cyc2, an outer membrane cytochrome c originally identified in Acidithiobacillus ferrooxidans. The "porin-cytochrome c complex" (PCC) gene clusters homologous to MtoAB/PioAB are present in eight FeOB, accounting for 19% of complete and close-to-complete genomes examined, whereas PCC genes homologous to OmbB-OmaB-OmcB in Geobacter sulfurreducens are absent. Further, we discovered gene clusters that may potentially encode two novel PCC types. First, a cluster (tentatively named "PCC3") encodes a porin, an extracellular and a periplasmic cytochrome c with remarkably large numbers of heme-binding motifs. Second, a cluster (tentatively named "PCC4") encodes a porin and three periplasmic multiheme cytochromes c. A conserved inner membrane protein (IMP) encoded in PCC3 and PCC4 gene clusters might be responsible for translocating electrons across the inner membrane. Other bacteria possessing PCC3 and PCC4 are mostly Proteobacteria isolated from environments with a potential niche for Fe(II) oxidation. In addition to cytochrome c, multicopper oxidase (MCO) genes potentially involved in Fe(II) oxidation were also identified. Notably, candidate EET genes were not found in some FeOB, especially the anaerobic ones, probably suggesting EET genes or Fe(II) oxidation mechanisms are different from the searched models. Overall, based on current EET models, the search extends our understanding of bacterial EET and provides candidate genes for future research.
Keywords: cytochrome c; extracellular electron transfer (EET); genomics; multicopper oxidase; neutrophilic Fe(II) oxidation; porin-cytochrome c complex (PCC).
Figures





References
-
- Barco R. A., Emerson D., Sylvan J. B., Orcutt B. N., Jacobson Meyers M. E., Ramirez G. A., et al. (2015). New insight into microbial iron oxidation as revealed by the proteomic profile of an obligate iron-oxidizing chemolithoautotroph. Appl. Environ. Microbiol. 81, 5927–5937. 10.1128/AEM.01374-15 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

