Toward the human cellular microRNAome
- PMID: 28877962
- PMCID: PMC5630040
- DOI: 10.1101/gr.222067.117
Toward the human cellular microRNAome
Abstract
MicroRNAs are short RNAs that serve as regulators of gene expression and are essential components of normal development as well as modulators of disease. MicroRNAs generally act cell-autonomously, and thus their localization to specific cell types is needed to guide our understanding of microRNA activity. Current tissue-level data have caused considerable confusion, and comprehensive cell-level data do not yet exist. Here, we establish the landscape of human cell-specific microRNA expression. This project evaluated 8 billion small RNA-seq reads from 46 primary cell types, 42 cancer or immortalized cell lines, and 26 tissues. It identified both specific and ubiquitous patterns of expression that strongly correlate with adjacent superenhancer activity. Analysis of unaligned RNA reads uncovered 207 unknown minor strand (passenger) microRNAs of known microRNA loci and 495 novel putative microRNA loci. Although cancer cell lines generally recapitulated the expression patterns of matched primary cells, their isomiR sequence families exhibited increased disorder, suggesting DROSHA- and DICER1-dependent microRNA processing variability. Cell-specific patterns of microRNA expression were used to de-convolute variable cellular composition of colon and adipose tissue samples, highlighting one use of these cell-specific microRNA expression data. Characterization of cellular microRNA expression across a wide variety of cell types provides a new understanding of this critical regulatory RNA species.
© 2017 McCall et al.; Published by Cold Spring Harbor Laboratory Press.
Figures
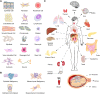



References
-
- Ambros V. 2004. The functions of animal microRNAs. Nature 431: 350–355. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
