HPV16 E7 Genetic Conservation Is Critical to Carcinogenesis
- PMID: 28886384
- PMCID: PMC5674785
- DOI: 10.1016/j.cell.2017.08.001
HPV16 E7 Genetic Conservation Is Critical to Carcinogenesis
Abstract
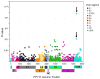
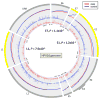
Although most cervical human papillomavirus type 16 (HPV16) infections become undetectable within 1-2 years, persistent HPV16 causes half of all cervical cancers. We used a novel HPV whole-genome sequencing technique to evaluate an exceptionally large collection of 5,570 HPV16-infected case-control samples to determine whether viral genetic variation influences risk of cervical precancer and cancer. We observed thousands of unique HPV16 genomes; very few women shared the identical HPV16 sequence, which should stimulate a careful re-evaluation of the clinical implications of HPV mutation rates, transmission, clearance, and persistence. In case-control analyses, HPV16 in the controls had significantly more amino acid changing variants throughout the genome. Strikingly, E7 was devoid of variants in precancers/cancers compared to higher levels in the controls; we confirmed this in cancers from around the world. Strict conservation of the 98 amino acids of E7, which disrupts Rb function, is critical for HPV16 carcinogenesis, presenting a highly specific target for etiologic and therapeutic research.
Keywords: E7 gene; HPV epidemiology; HPV genomics; HPV16; cervical carcinogenesis.
Published by Elsevier Inc.
Figures

Comment in
-
Crowd Control: E7 Conservation Is the Key to Cancer.Cell. 2017 Sep 7;170(6):1057-1059. doi: 10.1016/j.cell.2017.08.033. Cell. 2017. PMID: 28886377
References
-
- Bosch FX, Manos MM, Muñoz N, Sherman M, Jansen AM, Peto J, Schiffman MH, Moreno V, Kurman R, Shah KV. Prevalence of human papillomavirus in cervical cancer: a worldwide perspective. International biological study on cervical cancer (IBSCC) Study Group. J Natl Cancer Inst. 1995;87:796–802. - PubMed
-
- Burk RD, Ho GYF, Beardsley L, Lempa M, Peters M, Bierman R. Sexual behavior and partner characteristics are the predominant risk factors for genital human papillomavirus infection in young women. J Infect Dis. 1996;174:679–689. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

