Analysis of Class I Major Histocompatibility Complex Gene Transcription in Human Tumors Caused by Human Papillomavirus Infection
- PMID: 28891951
- PMCID: PMC5618018
- DOI: 10.3390/v9090252
Analysis of Class I Major Histocompatibility Complex Gene Transcription in Human Tumors Caused by Human Papillomavirus Infection
Abstract
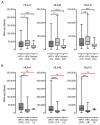
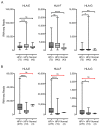
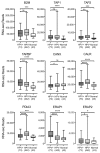
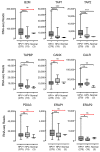
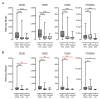
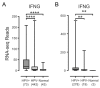
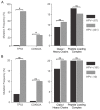
Oncoproteins from high-risk human papillomaviruses (HPV) downregulate the transcription of the class I major histocompatibility complex (MHC-I) antigen presentation apparatus in tissue culture model systems. This could allow infected or transformed cells to evade the adaptive immune response. Using data from over 800 human cervical and head & neck tumors from The Cancer Genome Atlas (TCGA), we determined the impact of HPV status on the mRNA expression of all six MHC-I heavy chain genes, and the β2 microglobulin light chain. Unexpectedly, these genes were all expressed at high levels in HPV positive (HPV+) cancers compared with normal control tissues. Indeed, many of these genes were expressed at significantly enhanced levels in HPV+ tumors. Similarly, the transcript levels of several other components of the MHC-I peptide-loading complex were also high in HPV+ cancers. The coordinated expression of high mRNA levels of the MHC-I antigen presentation apparatus could be a consequence of the higher intratumoral levels of interferon γ in HPV+ carcinomas, which correlate with signatures of increased infiltration by T- and NK-cells. These data, which were obtained from both cervical and oral tumors in large human cohorts, indicates that HPV oncoproteins do not efficiently suppress the transcription of the antigen presentation apparatus in human tumors.
Keywords: MHC-I; antigen presentation; cervical carcinoma; head & human papillomavirus; immune evasion; major histocompatibility complex; neck carcinoma.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

