Mitigating Mitochondrial Genome Erosion Without Recombination
- PMID: 28893855
- PMCID: PMC5676227
- DOI: 10.1534/genetics.117.300273
Mitigating Mitochondrial Genome Erosion Without Recombination
Abstract
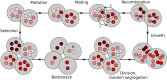
Mitochondria are ATP-producing organelles of bacterial ancestry that played a key role in the origin and early evolution of complex eukaryotic cells. Most modern eukaryotes transmit mitochondrial genes uniparentally, often without recombination among genetically divergent organelles. While this asymmetric inheritance maintains the efficacy of purifying selection at the level of the cell, the absence of recombination could also make the genome susceptible to Muller's ratchet. How mitochondria escape this irreversible defect accumulation is a fundamental unsolved question. Occasional paternal leakage could in principle promote recombination, but it would also compromise the purifying selection benefits of uniparental inheritance. We assess this tradeoff using a stochastic population-genetic model. In the absence of recombination, uniparental inheritance of freely-segregating genomes mitigates mutational erosion, while paternal leakage exacerbates the ratchet effect. Mitochondrial fusion-fission cycles ensure independent genome segregation, improving purifying selection. Paternal leakage provides opportunity for recombination to slow down the mutation accumulation, but always at a cost of increased steady-state mutation load. Our findings indicate that random segregation of mitochondrial genomes under uniparental inheritance can effectively combat the mutational meltdown, and that homologous recombination under paternal leakage might not be needed.
Keywords: Muller’s ratchet; maternal inheritance; mitochondrial recombination; paternal leakage; uniparental inheritance.
Copyright © 2017 by the Genetics Society of America.
Figures



References
-
- Burger G., Gray M. W., Lang B. F., 2003. Mitochondrial genomes: anything goes. Trends Genet. 19: 709–716. - PubMed
-
- Buss L. W., 1987. The Evolution of Individuality. Princeton University Press, Princeton, NJ.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

