Nanobodies: Chemical Functionalization Strategies and Intracellular Applications
- PMID: 28913971
- PMCID: PMC5838514
- DOI: 10.1002/anie.201708459
Nanobodies: Chemical Functionalization Strategies and Intracellular Applications
Abstract
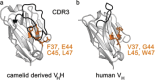
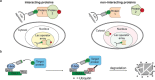
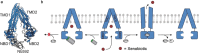
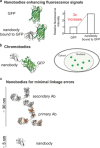
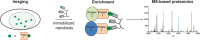
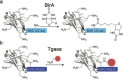
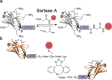
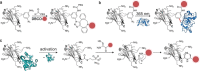
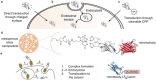
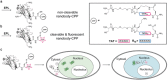
Nanobodies can be seen as next-generation tools for the recognition and modulation of antigens that are inaccessible to conventional antibodies. Due to their compact structure and high stability, nanobodies see frequent usage in basic research, and their chemical functionalization opens the way towards promising diagnostic and therapeutic applications. In this Review, central aspects of nanobody functionalization are presented, together with selected applications. While early conjugation strategies relied on the random modification of natural amino acids, more recent studies have focused on the site-specific attachment of functional moieties. Such techniques include chemoenzymatic approaches, expressed protein ligation, and amber suppression in combination with bioorthogonal modification strategies. Recent applications range from sophisticated imaging and mass spectrometry to the delivery of nanobodies into living cells for the visualization and manipulation of intracellular antigens.
Keywords: antigen-binding proteins; cellular delivery; molecular biology; nanobodies; site-specific functionalization.
© 2017 The Authors. Published by Wiley-VCH Verlag GmbH & Co. KGaA.
Figures





References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

