The Effect of Common Inversion Polymorphisms In(2L)t and In(3R)Mo on Patterns of Transcriptional Variation in Drosophila melanogaster
- PMID: 28916647
- PMCID: PMC5677173
- DOI: 10.1534/g3.117.1133
The Effect of Common Inversion Polymorphisms In(2L)t and In(3R)Mo on Patterns of Transcriptional Variation in Drosophila melanogaster
Abstract
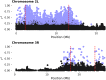
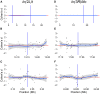
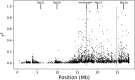
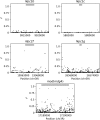
Chromosomal inversions are a ubiquitous feature of genetic variation. Theoretical models describe several mechanisms by which inversions can drive adaptation and be maintained as polymorphisms. While inversions have been shown previously to be under selection, or contain genetic variation under selection, the specific phenotypic consequences of inversions leading to their maintenance remain unclear. Here we use genomic sequence and expression data from the Drosophila Genetic Reference Panel (DGRP) to explore the effects of two cosmopolitan inversions, In(2L)t and In(3R)Mo, on patterns of transcriptional variation. We demonstrate that each inversion has a significant effect on transcript abundance for hundreds of genes across the genome. Inversion-affected loci (IAL) appear both within inversions as well as on unlinked chromosomes. Importantly, IAL do not appear to be influenced by the previously reported genome-wide expression correlation structure. We found that five genes involved with sterol uptake, four of which are Niemann-Pick Type 2 orthologs, are upregulated in flies with In(3R)Mo but do not have SNPs in linkage disequilibrium (LD) with the inversion. We speculate that this upregulation is driven by genetic variation in mod(mdg4) that is in LD with In(3R)Mo We find that there is little evidence for a regional or position effect of inversions on gene expression at the chromosomal level, but do find evidence for the distal breakpoint of In(3R)Mo interrupting one gene and possibly disassociating the two flanking genes from regulatory elements.
Keywords: DGRP; DPGP; Drosophila; eQTLs; inversion polymorphism.
Copyright © 2017 Lavington and Kern.
Figures

Similar articles
-
Genomic Evidence for Adaptive Inversion Clines in Drosophila melanogaster.Mol Biol Evol. 2016 May;33(5):1317-36. doi: 10.1093/molbev/msw016. Epub 2016 Jan 21. Mol Biol Evol. 2016. PMID: 26796550
-
Patterns of diversity and linkage disequilibrium within the cosmopolitan inversion In(3R)Payne in Drosophila melanogaster are indicative of coadaptation.Genetics. 2006 Mar;172(3):1655-63. doi: 10.1534/genetics.105.053173. Epub 2005 Dec 1. Genetics. 2006. PMID: 16322502 Free PMC article.
-
Genomic evidence for role of inversion 3RP of Drosophila melanogaster in facilitating climate change adaptation.Mol Ecol. 2015 May;24(10):2423-32. doi: 10.1111/mec.13161. Epub 2015 Apr 20. Mol Ecol. 2015. PMID: 25789416
-
Patterns of inversion polymorphism in three species of the Drosophila melanogaster species group.Indian J Exp Biol. 2001 Jul;39(7):611-22. Indian J Exp Biol. 2001. PMID: 12019752 Review.
-
Population genetics of inversion polymorphism in Drosophila ananassae.Indian J Exp Biol. 1998 Aug;36(8):739-48. Indian J Exp Biol. 1998. PMID: 9838874 Review.
Cited by
-
Association mapping desiccation resistance within chromosomal inversions in the African malaria vector Anopheles gambiae.Mol Ecol. 2019 Mar;28(6):1333-1342. doi: 10.1111/mec.14880. Epub 2018 Oct 22. Mol Ecol. 2019. PMID: 30252170 Free PMC article.
-
Genetic architecture of repeated phenotypic divergence in Littorina saxatilis ecotype evolution.Evolution. 2022 Oct;76(10):2332-2346. doi: 10.1111/evo.14602. Epub 2022 Sep 1. Evolution. 2022. PMID: 35994296 Free PMC article.
-
A large chromosomal inversion shapes gene expression in seaweed flies (Coelopa frigida).Evol Lett. 2021 Oct 7;5(6):607-624. doi: 10.1002/evl3.260. eCollection 2021 Dec. Evol Lett. 2021. PMID: 34917400 Free PMC article.
-
The Role of Structural Variation in Adaptation and Evolution of Yeast and Other Fungi.Genes (Basel). 2021 May 8;12(5):699. doi: 10.3390/genes12050699. Genes (Basel). 2021. PMID: 34066718 Free PMC article. Review.
-
How chromosomal rearrangements shape adaptation and speciation: Case studies in Drosophila pseudoobscura and its sibling species Drosophila persimilis.Mol Ecol. 2019 Mar;28(6):1283-1301. doi: 10.1111/mec.14923. Epub 2018 Dec 10. Mol Ecol. 2019. PMID: 30402909 Free PMC article.
References
-
- Anderson A. R., Hoffmann A. A., McKechnie S. W., Umina P. A., Weeks A. R., 2005. The latitudinal cline in the In(3R)Payne inversion polymorphism has shifted in the last 20 years in Australian Drosophila melanogaster populations. Mol. Ecol. 14: 851–858. - PubMed
-
- Anderson W. W., Dobzhansky T., Kastritsis C. D., 1967. Selection and inversion polymorphism in experimental populations of Drosophila pseudoobscura initiated with the chromosomal constitutions of natural populations. Evolution 21: 664–671. - PubMed
-
- Andolfatto P., Depaulis F., Navarro A., 2001. Inversion polymorphisms and nucleotide variability in Drosophila. Genet. Res. 77: 1–8. - PubMed
-
- Aulard S., David J. R., Lemeunier F., 2002. Chromosomal inversion polymorphism in Arotropical populations of Drosophila melanogaster. Genet. Res. 79: 49–63. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
