CRISPR/Cas9 Engineering of Adult Mouse Liver Demonstrates That the Dnajb1-Prkaca Gene Fusion Is Sufficient to Induce Tumors Resembling Fibrolamellar Hepatocellular Carcinoma
- PMID: 28923495
- PMCID: PMC5801691
- DOI: 10.1053/j.gastro.2017.09.008
CRISPR/Cas9 Engineering of Adult Mouse Liver Demonstrates That the Dnajb1-Prkaca Gene Fusion Is Sufficient to Induce Tumors Resembling Fibrolamellar Hepatocellular Carcinoma
Abstract
Background & aims: Fibrolamellar hepatocellular carcinoma (FL-HCC) is a primary liver cancer that predominantly affects children and young adults with no underlying liver disease. A somatic, 400 Kb deletion on chromosome 19 that fuses part of the DnaJ heat shock protein family (Hsp40) member B1 gene (DNAJB1) to the protein kinase cAMP-activated catalytic subunit alpha gene (PRKACA) has been repeatedly identified in patients with FL-HCC. However, the DNAJB1-PRKACA gene fusion has not been shown to induce liver tumorigenesis. We used the CRISPR/Cas9 technique to delete in mice the syntenic region on chromosome 8 to create a Dnajb1-Prkaca fusion and monitored the mice for liver tumor development.
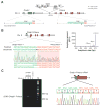
Methods: We delivered CRISPR/Cas9 vectors designed to juxtapose exon 1 of Dnajb1 with exon 2 of Prkaca to create the Dnajb1-Prkaca gene fusion associated with FL-HCC, or control Cas9 vector, via hydrodynamic tail vein injection to livers of 8-week-old female FVB/N mice. These mice did not have any other engineered genetic alterations and were not exposed to liver toxins or carcinogens. Liver tissues were collected 14 months after delivery; genomic DNA was analyzed by PCR to detect the Dnajb1-Prkaca fusion, and tissues were characterized by histology, immunohistochemistry, RNA sequencing, and whole-exome sequencing.
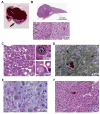
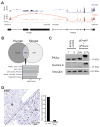
Results: Livers from 12 of the 15 mice given the vectors to induce the Dnajb1-Prkaca gene fusion, but none of the 11 mice given the control vector, developed neoplasms. The tumors contained the Dnajb1-Prkaca gene fusion and had histologic and cytologic features of human FL-HCCs: large polygonal cells with granular, eosinophilic, and mitochondria-rich cytoplasm, prominent nucleoli, and markers of hepatocytes and cholangiocytes. In comparing expression levels of genes between the mouse tumor and non-tumor liver cells, we identified changes similar to those detected in human FL-HCC, which included genes that affect cell cycle and mitosis regulation. Genomic analysis of mouse neoplasms induced by the Dnajb1-Prkaca fusion revealed a lack of mutations in genes commonly associated with liver cancers, as observed in human FL-HCC.
Conclusions: Using CRISPR/Cas9 technology, we found generation of the Dnajb1-Prkaca fusion gene in wild-type mice to be sufficient to initiate formation of tumors that have many features of human FL-HCC. Strategies to block DNAJB1-PRKACA might be developed as therapeutics for this form of liver cancer.
Keywords: Genomic Engineering; Liver Cancer; Mouse Model; PKA; Protein Kinase A.
Copyright © 2017 AGA Institute. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Figures

Similar articles
-
Fibrolamellar Carcinoma: Recent Advances and Unresolved Questions on the Molecular Mechanisms.Semin Liver Dis. 2018 Feb;38(1):51-59. doi: 10.1055/s-0037-1621710. Epub 2018 Feb 22. Semin Liver Dis. 2018. PMID: 29471565 Free PMC article. Review.
-
DNAJB1-PRKACA fusion kinase interacts with β-catenin and the liver regenerative response to drive fibrolamellar hepatocellular carcinoma.Proc Natl Acad Sci U S A. 2017 Dec 12;114(50):13076-13084. doi: 10.1073/pnas.1716483114. Epub 2017 Nov 21. Proc Natl Acad Sci U S A. 2017. PMID: 29162699 Free PMC article.
-
DNAJB1-PRKACA fusions occur in oncocytic pancreatic and biliary neoplasms and are not specific for fibrolamellar hepatocellular carcinoma.Mod Pathol. 2020 Apr;33(4):648-656. doi: 10.1038/s41379-019-0398-2. Epub 2019 Nov 1. Mod Pathol. 2020. PMID: 31676785 Free PMC article.
-
A genomic case study of mixed fibrolamellar hepatocellular carcinoma.Ann Oncol. 2016 Jun;27(6):1148-1154. doi: 10.1093/annonc/mdw135. Epub 2016 Mar 30. Ann Oncol. 2016. PMID: 27029710 Free PMC article.
-
Fibrolamellar Hepatocellular Carcinoma: Mechanistic Distinction From Adult Hepatocellular Carcinoma.Pediatr Blood Cancer. 2016 Jul;63(7):1163-7. doi: 10.1002/pbc.25970. Epub 2016 Mar 14. Pediatr Blood Cancer. 2016. PMID: 26990031 Free PMC article. Review.
Cited by
-
Fibrolamellar hepatocellular carcinoma: A rare but unpleasant event.World J Gastrointest Oncol. 2022 Jun 15;14(6):1103-1114. doi: 10.4251/wjgo.v14.i6.1103. World J Gastrointest Oncol. 2022. PMID: 35949219 Free PMC article. Review.
-
Fibrolamellar Carcinoma: Recent Advances and Unresolved Questions on the Molecular Mechanisms.Semin Liver Dis. 2018 Feb;38(1):51-59. doi: 10.1055/s-0037-1621710. Epub 2018 Feb 22. Semin Liver Dis. 2018. PMID: 29471565 Free PMC article. Review.
-
An acquired scaffolding function of the DNAJ-PKAc fusion contributes to oncogenic signaling in fibrolamellar carcinoma.Elife. 2019 May 7;8:e44187. doi: 10.7554/eLife.44187. Elife. 2019. PMID: 31063128 Free PMC article.
-
Increased Protein Kinase A Activity Induces Fibrolamellar Hepatocellular Carcinoma Features Independent of DNAJB1.Cancer Res. 2024 Aug 15;84(16):2626-2644. doi: 10.1158/0008-5472.CAN-23-4110. Cancer Res. 2024. PMID: 38888469 Free PMC article.
-
Liver-directed therapies for fibrolamellar carcinoma: A single-center experience.Oncol Res. 2024 Nov 13;32(12):1831-1836. doi: 10.32604/or.2024.052985. eCollection 2024. Oncol Res. 2024. PMID: 39574471 Free PMC article.
References
-
- Kakar S, Burgart LJ, Batts KP, et al. Clinicopathologic features and survival in fibrolamellar carcinoma: comparison with conventional hepatocellular carcinoma with and without cirrhosis. Mod Pathol. 2005;18(11):1417–23. - PubMed
-
- Craig JR, Peters RL, Edmondson HA, et al. Fibrolamellar carcinoma of the liver: a tumor of adolescents and young adults with distinctive clinico-pathologic features. Cancer. 1980;46(2):372–9. - PubMed
-
- El-Serag HB, Davila JA. Is fibrolamellar carcinoma different from hepatocellular carcinoma? A US population-based study. Hepatology. 2004;39(3):798–803. - PubMed
-
- Ward SC, Huang J, Tickoo SK, et al. Fibrolamellar carcinoma of the liver exhibits immunohistochemical evidence of both hepatocyte and bile duct differentiation. Mod Pathol. 2010;23(9):1180–90. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

