A vital sugar code for ricin toxicity
- PMID: 28925387
- PMCID: PMC5674155
- DOI: 10.1038/cr.2017.116
A vital sugar code for ricin toxicity
Abstract
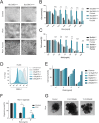
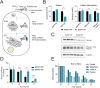
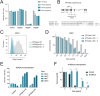
Ricin is one of the most feared bioweapons in the world due to its extreme toxicity and easy access. Since no antidote exists, it is of paramount importance to identify the pathways underlying ricin toxicity. Here, we demonstrate that the Golgi GDP-fucose transporter Slc35c1 and fucosyltransferase Fut9 are key regulators of ricin toxicity. Genetic and pharmacological inhibition of fucosylation renders diverse cell types resistant to ricin via deregulated intracellular trafficking. Importantly, cells from a patient with SLC35C1 deficiency are also resistant to ricin. Mechanistically, we confirm that reduced fucosylation leads to increased sialylation of Lewis X structures and thus masking of ricin-binding sites. Inactivation of the sialyltransferase responsible for modifications of Lewis X (St3Gal4) increases the sensitivity of cells to ricin, whereas its overexpression renders cells more resistant to the toxin. Thus, we have provided unprecedented insights into an evolutionary conserved modular sugar code that can be manipulated to control ricin toxicity.
Figures



References
-
- Audi J, Belson M, Patel M, Schier J, Osterloh J. Ricin poisoning: a comprehensive review. JAMA 2005; 294:2342–2351. - PubMed
-
- Sandvig K, Skotland T, van Deurs B, Klokk TI. Retrograde transport of protein toxins through the Golgi apparatus. Histochem Cell Biol 2013; 140:317–326. - PubMed
-
- Crompton R, Gall D. Georgi Markov — death in a pellet. Med Leg J 1980; 48: 51–62. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

